Patent application title: AIB1, a novel steroid receptor co-activator
Inventors:
Paul Meltzer (Rockville, MD, US)
Jeffrey Trent (Rockville, MD, US)
Assignees:
The Government of the United States of America as Represented by the Secretary
of the Department of Health and Human Services
IPC8 Class: AA61K3817FI
USPC Class:
514 12
Class name: Designated organic active ingredient containing (doai) peptide containing (e.g., protein, peptones, fibrinogen, etc.) doai 25 or more peptide repeating units in known peptide chain structure
Publication date: 2009-01-22
Patent application number: 20090023645
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: AIB1, a novel steroid receptor co-activator
Inventors:
Paul Meltzer
Jeffrey Trent
Agents:
KLARQUIST SPARKMAN, LLP
Assignees:
Origin: PORTLAND, OR US
IPC8 Class: AA61K3817FI
USPC Class:
514 12
Abstract:
The invention features a substantially pure DNA which includes a sequence
encoding a novel steroid receptor co-activator which is overexpressed in
breast cancer cells, diagnostic assays for steroid hormone-responsive
cancers, and screening assays to identify compounds which inhibit an
interaction of the co-activator with the steroid hormone.Claims:
1. A method of screening a candidate compound which inhibits an
interaction of an AIB1 polypeptide with an ER polypeptide in a cell
comprising(a) providing a GAL4 binding site linked to a reporter gene;(b)
providing a GAL4 binding domain linked to either (i) an AIB1 polypeptide
or (ii) an ER polypeptide;(c) providing a GAL4 transactivation domain II
linked to the ER polypeptide if the GAL4 binding domain is linked to the
AIB1 polypeptide or linked to the AIB1 polypeptide if the GAL4 binding
domain is linked to the ER polypeptide;(d) contacting the cell with the
compound; and(e) monitoring expression of the reporter gene, wherein a
decrease in expression in the presence of the compound compared to that
in the absence of the compound indicates that the compound inhibits an
interaction of an AIB1 polypeptide with the ER polypeptide.
2. A method of detecting an aberrantly proliferating cell in a tissue sample comprising determining the level of AIB1 gene expression in the sample, wherein an increase in the level of expression compared to the level in normal control tissue indicates the presence of an aberrantly proliferating cell.
3. The method of claim 2, wherein the aberrantly proliferating cell is a steroid hormone-responsive cancer cell.
4. The method of claim 3, wherein the steroid hormone-responsive cancer cell is a breast cancer cell.
5. The method of claim 3, wherein the cell is a steroid hormone-responsive cancer cell is an ovarian cancer cell.
6. The method of claim 1, wherein the AIB1 gene expression is measured using an AIB1 gene-specific polynucleotide probe.
7. The method of claim 1, wherein the AIB1 gene expression is measured using an antibody that specifically binds AIB1.
8. A method of detecting breast cancer in a tissue sample, comprising determining the number of cellular copies of an AIB1 gene in the tissue sample, wherein an increase in the number of copies compared to the number of copies in a normal control tissue indicates the presence of a breast carcinoma.
9. The method of claim 8, wherein the number of copies in the tissue is greater than 2.
10. The method of claim 9, wherein the number of copies in the tissue is greater than 10.
11. The method of claim 10, wherein the number of copies in the tissue is greater than 20.
12. A method of reducing proliferation of a cancer cell in a mammal comprising administering to the mammal a compound which inhibits expression of AIB1.
13. The method of claim 12, wherein the compound reduces transcription of DNA encoding AIB1 in the cell.
14. The method of claim 12, wherein the compound reduces translation of an AIB1 mRNA into an AIB1 gene product in the cell.
15. The method of claim 14, wherein the translation is reduced by contacting the AIB1 mRNA with an antisense DNA complementary to the AIB1 mRNA.
16. A method of inhibiting ER-dependent transcription in a breast cell of an mammal, comprising administering an effective amount of an AIB1 polypeptide to the mammal.
17. The method of claim 16, wherein the polypeptide comprises a PAS domain.
18. The method of claim 16, wherein the polypeptide comprises a bHLH domain.
19. The method of claim 16, wherein the polypeptide comprises an ER-interacting domain
20. A method of identifying a tamoxifen-sensitive patient, comprising(a) contacting a patient-derived tissue sample with tamoxifen; and(b) determining the level of AIB1 gene expression in the sample, wherein an increase in the level of expression compared to the level in normal control tissue indicates that the patient is tamoxifen-sensitive.
21. The method of claim 20, wherein the AIB1 gene expression is measured using an AIB1 gene-specific polynucleotide probe.
22. The method of claim 20, wherein the AIB1 gene expression is measured using an antibody that specifically binds AIB1.
23. A method of reducing proliferation of a cancer cell comprising administering to the mammal a compound which inhibits interaction of AIB1 with a molecule selected from the group consisting of steroid receptors and nuclear co-factors.
24. The method of claim 23 wherein the molecule is p300 or CBP.
Description:
PRIORITY CLAIM
[0001]This is a divisional of U.S. patent application Ser. No. 10/379,616, filed Mar. 4, 2003, incorporated by reference herein. U.S. patent application Ser. No. 10/379,616 is a divisional of U.S. patent application Ser. No. 09/125,635, filed Aug. 21, 1998, now issued as U.S. Pat. No. 6,562,589. U.S. patent application Ser. No. 09/125,635 is a §371 U.S. national stage of PCT/US98/12689, filed Jun. 17, 1998, which was published in English under PCT Article 21(2), which claims the benefit of U.S. Provisional Application No. 60/049,728, filed Jun. 17, 1997.
BACKGROUND OF THE INVENTION
[0002]Breast cancer arises from estrogen-responsive breast epithelial cells. Estrogen activity is thought to promote the development of breast cancer, and many breast cancers are initially dependent on estrogen at the time of diagnosis. Anti-estrogen compositions have therefore been used to treat breast cancer.
[0003]A frequent mechanism of increased gene expression in human cancers is amplification, i.e., the copy number of a DNA sequence is increased, in a cancer cell compared to a non-cancerous cell. In breast cancer, commonly amplified regions are derived from 17q21, 8q24, and 11q13 which encode erbB-2, c-myc, and cyclic DI respectively (Devilee et al., 1994, Crit. Rev. Oncog. 5:247-270). Recently, molecular cytogenetic studies have revealed the occurrence in breast cancers of additional regions of increased DNA copy number (Isola et al., Am. J. Pathol. 147:905-911, 1995; Kallioniemi et al., Proc. Natl. Acad. Sci. USA 91:2156-2160, 1994; Muleris et al., Genes Chromo. Cancer 10:160-170, 1994; Tanner et al., Cancer Research 54:4257-4260, 1994; Guan et al., Nat. Genet. 8:155-161, 1994).
[0004]Breast cancer is the second leading cause of cancer deaths in American women, and it is estimated that an American woman has at least a 10% cumulative lifetime risk of developing this disease. Early diagnosis is an important factor in breast cancer prognosis and affects not only survival rate, but the range of therapeutic options available to the patient. For instance, if diagnosed early, a "lumpectomy" may be performed, whereas later diagnosis tends to be associated with more invasive and traumatic surgical treatments such as radical mastectomy. The treatment of other cancers likewise is benefitted by early diagnosis, for instance the prognosis in the treatment of lung cancer, colorectal cancer and prostate cancers is greatly improved by early diagnosis. There is a need for a simple and reliable method of diagnosis of cancers in general and of breast cancer in particular. There is a need for a method of screening for compounds that inhibit the interaction between an estrogen receptor ER and an ER-dependent nuclear receptor co-activator molecule in order to identify molecules useful in research diagnosis and treatment of cancer. There is also a need for a method for identifying tamoxifen-sensitive cancer patients in order to better manage treatment. A solution to these needs would improve cancer treatment and research and would save lives.
SUMMARY OF THE INVENTION
[0005]The inventors have discovered that the AIB1 protein (Amplified In Breast Cancer-1) is a member of the Steroid Receptor Coactivator-1 (SRC-1) family of nuclear receptor co-activators that interacts with estrogen receptors (ER) to enhance ER-dependent transcription. The inventors have further discovered that the AIB1 gene is amplified and over-expressed in certain cancers including breast cancer, and that detection of amplified AIB1 genes can therefore be used to detect cancerous cells. Importantly, the inventors have also found that AIB1 amplification is not confined to breast cancer but is also found in cancers of the lung, ovary, head and neck, colon, testicles, bladder, prostate, endometrium, kidney, stomach and also in pheochromocytoma, melanoma, ductal carcinoma and carcinoid tumor. Such a finding means that AIB1 may be useful in the detection and treatment of all of the aforementioned cancers which include some of the most prevalent and deadly diseases in the western world.
[0006]The inventors have also discovered that AIB1 interacts with the proteins p300 and CBP, which are nuclear cofactors that interact with other nuclear factors to promote transcription (Chacravarti et al., Nature (383) 99-103 1996; Lundblad et al., Nature (374) 85-88 1995). The inventors have, furthermore, determined that in cells with stable over-expression of AIB1, there is a dramatic increase in steroid receptor activation (almost a 100-fold increase) leading to a corresponding increase in transcriptional activation. The inventors have also used monoclonal anti-AIB1 antibodies to demonstrate that AIB1 gene amplification is directly correlated with increased AIB1 expression, and that these amplified copies of the gene are expressed in physiological conditions. The inventors have found that AIB1 is the human ortholog of the mouse ER-dependent transcriptional activator p/CIP, with the proteins having an overall amino acid identity of 81.6%. These finding support the physiological role for AIB1 in cancer cells as a cofactor involved in transcriptional regulation.
[0007]The invention features a substantially pure DNA which includes a sequence encoding an AIB1 polypeptide, e.g., a human AIB1 polypeptide, or a fragment thereof. The DNA may have the sequence of all or part of the naturally-occurring AIB1-encoding DNA or a degenerate variant thereof. AIB1-encoding DNA may be operably linked to regulatory sequences for expression of the polypeptide. A cell containing AIB1 encoding DNA is also within the invention.
[0008]The invention also includes a substantially pure DNA containing a polynucleotides which hybridizes at high stringency to a AIB1-encoding DNA or the complement thereof. A substantially pure DNA containing a nucleotide sequence having at least 50% sequence identity to the full length AIB1 cDNA, e.g., a nucleotide sequence encoding a polypeptide having the biological activity of a AIB1 polypeptide, is also included.
[0009]The invention also features a substantially pure human AIB1 polypeptide and variants thereof, e.g., polypeptides with conservative amino acid substitutions or polypeptides with conservative or non-conservative amino acid substitutions which retain the biological activity of naturally-occurring AIB1.
[0010]Diagnostic methods, e.g., to identify cells which harbor an abnormal copy number of the AIB1 DNA, are also encompassed by the invention. An abnormal copy number, e.g., greater than the normal diploid copy number, of AIB1 DNA is indicative of an aberrantly proliferating cell, e.g., a steroid hormone-responsive cancer cell.
[0011]The invention also includes antibodies, e.g., a monoclonal antibody or polyclonal antisera, which bind specifically to AIB1 and can be used to detect the level of expression of AIB1 in a cell or tissue sample. An increase in the level of expression of AIB1 in a patient-derived tissue sample compared to the level in normal control tissue indicates the presence of a cell proliferative disorder such as cancer.
[0012]Screening methods to identify compounds which inhibit an interaction of AIB1 with a steroid hormone receptor, thus disrupting a signal transduction pathway which leads to aberrant cell proliferation, is also within the invention. Proliferation of a cancer cell can therefore be reduced by administering to an individual, e.g., a patient diagnosed with a steroid-responsive cancer, a compound which inhibits expression of AIB1.
[0013]The invention also includes a knockout mutant, for example a mouse (or other mammal) from which at least one AIB1 gene has been selectively deleted from its genome. Such a mouse is useful in research, for instance, the phenotype gives insight into the physiological role of the deleted gene. For instance the mutant may be defective in specific biochemical pathways; such a knockout mutant may be used in complementation experiments to determine the role of other genes and proteins to determine if any such genes or proteins complement for the deleted gene. Homozygous and heterozygous mutants are included in this aspect of the invention.
[0014]The present invention also includes a mutant organism, for example a mammal such as a mouse which contains more than the normal number of AIB1 genes in its genome. Such a mouse may contain additional copies of the AIB1 gene integrated into its chromosomes, for instance in the form of a pro-virus, or may carry additional copies on extra-chromosomal elements such as plasmids. Such a mutant mouse is useful for research purposes, to elucidate the physiological or pathological role of AIB1. For instance, the role of AIB1 expression as cause or effect in cancers may be investigated by including or transplanting tumors into such mutants, and comparing such mutants with normal mice having the same cancer.
[0015]The present invention also includes a mutant organism, for example a mammal, e.g. a mouse, that contains, either integrated into a chromosome or on a plasmid, at least one copy of the AIB1 gene driven by a non-native promoter. Such a promoter may be constitutive or may be inducible. For instance, the AIB1 gene may be operatively linked to a mouse mammary tumor virus (MMTV) promoter or other promoter from a mammalian virus allowing manipulation of AIB1 expression. Such a mutant would be useful for research purposes to determine the physiological or pathological role of AIB1. For instance, over or under expression could be affected and physiological effects observed.
[0016]The invention also includes methods for treatment of cancers that involve functions of or alterations in the signaling pathways that use p300 and/or CBP as signal transducing molecules. The treatments of the invention involve targeting of the AIB1 protein or AIB1 gene to enhance or reduce interaction with p300 and/or CBP proteins. For instance, the AIB1 gene sequence as disclosed herein may be used to construct an anti-sense nucleotide. An anti-sense RNA may be constructed that is anti-parallel and complementary to the AIB1 transcript (or part thereof) and which will therefore form an RNA-RNA duplex with the AIB1 transcript, preventing transcription and expression of AIB1. Alternatively, treatments may comprise contacting an AIB1 protein with a molecule that specifically binds to the AIB1 molecule in vivo, thereby interfering with AIB1 binding with other factors such as p300 or CBP. Such processes are designed to inhibit signal transduction pathways involving AIB1, p300, CBP and other factors and therefore inhibit cancer cell proliferation that is effected via these pathways. As explained in more detail below, AIB1 overexpression results in increased ER-dependent transcriptional activity which confers a growth advantage upon AIB1 amplification-bearing clones during the development and progression of estrogen-dependent cancers.
[0017]Compounds which inhibit or disrupt the interaction of an AIB1 gene product with a steroid hormone receptor, e.g., ER, are useful as anti-neoplastic agents for the treatment of patients suffering from steroid hormone-responsive cancers such as breast cancer, ovarian cancer, prostate cancer, and colon cancer.
[0018]AIB1 polypeptides or peptide mimetics of such polypeptides, e.g., those containing domains which interact with steroid hormone receptors, can be administered to patients to block the interaction of endogenous intracellular AIB1 and a steroid hormone receptor, e.g., ER in an aberrantly proliferating cell. It is likely that AIB1 interacts with a wide range of human transcriptional factors and that regulation of such interactions will have important therapeutic applications.
[0019]Other features and advantages of the invention will be apparent from the following description of the preferred embodiments thereof, and from the claims.
Sequence Listing
[0020]The nucleic acid and amino acid sequences listed in the accompanying Sequence Listing are shown using standard letter abbreviations for nucleotide bases and three-letter code for amino acids. Only one strand of each nucleic acid sequence is shown, but the complementary strand is understood to be included by any reference to the displayed strand.
[0021]SEQ ID NO: 1 shows the nucleic acid sequence of the human AIB1 cDNA.
[0022]SEQ ID NO: 2 shows the amino acid sequence of the Per/Arnt/Sim (PAS) domain of AIB1.
[0023]SEQ ID NO: 3 shows the amino acid sequence of the basic helix-loop-helix domain (bHLH) of AIB1.
[0024]SEQ ID NO: 4 shows the amino acid sequence of the human AIB1 protein.
[0025]SEQ ID NO: 5 shows the nucleic acid sequence of primer N8F1.
[0026]SEQ ID NO: 6 shows the nucleic acid sequence of the forward primer designed from the 5' sequence of pCMVSPORT-B11, PM-U2.
[0027]SEQ ID NO: 7 shows the nucleic acid sequence of the reverse primer designed from the 5' sequence of pCMVSPORT-B11, PM-U2.
[0028]SEQ ID NO: 8 shows the amino acid sequence of the ER-interacting domain of AIB1.
[0029]SEQ ID NO: 9 shows the nucleic acid sequence of pCIP, the mouse ortholog of AIB1 and the amino acid sequence for this gene.
[0030]SEQ ID NO: 10 shows the nucleic acid sequence of the forward primer AIB1/mESTF1 used to screen mouse BAC.
[0031]SEQ ID NO: 11 shows the nucleic acid sequence of the reverse primer AIB1/mESTR1 used to screen mouse BAC.
[0032]SEQ ID NO: 12 shows the amino acid sequence of pCIP, the mouse ortholog of AIB1.
BRIEF DESCRIPTION OF THE DRAWINGS
[0033]FIG. 1A is a diagram of an amino acid sequence of full length AIB1 (SEQ ID NO:4) in which residues highlighted in black are identical in AIB1, TIF2, and SRC1. Residues identical with TIF2 (GenBank Accession No. X97674) or SRC-1 (GenBank Accession No. U59302) are highlighted in grey or boxed, respectively.
[0034]FIG. 1B is a diagram showing the structural features of AIB1. The following domains are indicated: bHLH domain, PAS domains (with the highly conserved PAS A and B regions shown in dark gray), S/T (serine/threonine)-rich regions, and a group of charged residues (+/-). A glutamine-rich region and polyglutamine tract are also indicated. The numbers beneath the diagram indicate the location (approximate residue number) of the domain with respect to the amino acid sequence shown in FIG. 1A. The alignment was generated using DNASTAR software.
[0035]FIG. 2 is a photograph of a Northern blot analysis showing increased expression of AIB1 in the cell lines BT-474, ZR-75-1, MCF7, and BG-1.
[0036]FIG. 3 is a bar graph showing that the addition of full length AIB1 DNA to a cell resulted in an increase of estrogen-dependent transcription from an ER reporter plasmid. COS-1 cells were transiently transfected with 250 ng ER expression vector (pHEGO-hyg), 10 ng of luciferase reporter plasmid (pGL3.luc.3ERE or 10 ng pGL3 lacking ERE) and increasing amounts of pcDNA3.1-AIB1 and incubated in the absence (open bars) or presence of 10 nM 17β-stradiol (E2, solid bars) or 100 nM 4-hydroxytamoxifen (hatched bars). Luciferase activity was expressed in relative luminescence units (RLU). The data are the mean of three determinations from one of four replicate experiments. Error bars indicate one standard deviation.
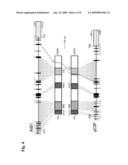
[0037]FIG. 4 is a schematic diagram comparing the DNA and protein structures of pCIP (the mouse ortholog of AIB1) and the human AIB1; exons are shown as black boxes.
[0038]FIGS. 5A and-5B are a table showing the introns and exons of the mouse AIB1 gene (pCIP) (SEQ ID NO:9). The "Exon" column refers to the number of the exon; "cDNA bp 5'-exon" refers to the nucleotide position in the mouse cDNA sequence for the 5' exon; "cDNA bp 3' exon" refers to the last few nucleotides of the 3' position of the intron. "Exon sequence" refers to the exon itself. "5' intron" refers to the adjacent intron reading from the exon into the splice donor nucleotides (usually GT).
[0039]FIG. 6A and FIG. 6B are a table showing the introns and exons of the human AIB1 gene (SEQ ID NO:1). The "Exon" column refers to the number of the exon; "cDNA bp 5'-exon" refers to the nucleotide position in the mouse cDNA sequence for the 5' exon; "cDNA bp 3' exon" refers to the last few nucleotides of the 3' position of the intron. "Exon sequence" refers to the exon itself. "5' intron" refers to the adjacent intron reading from the exon into the splice donor nucleotides (usually GT).
DETAILED DESCRIPTION
[0040]The invention is based on the discovery of a novel gene, amplified in breast cancer-1 (AIB1), which is overexpressed in breast cancer. AIB1 has the structural features of a co-activator of the steroid hormone receptor family. The steroid hormone estrogen and other related steroid hormones act on cells through specific steroid receptors.
[0041]Members of the steroid receptor coactivator (SRC) family of transcriptional co-activators interact with nuclear hormone receptors to enhance ligand-dependent transcription. AIB1 is a novel member of the SRC family which was found to be overexpressed in breast cancers. The AIB1 gene is located at human chromosome 20q. High-level AIB1 amplification and overexpression were observed in several estrogen receptor (ER) positive breast and ovarian cancer cell lines, as well as in uncultured breast cancer specimens. AIB1 amplification is not confined to breast cancer but is also found in cancers of the lung, ovary, head and neck, colon, testicles, bladder, prostate, endometrium, kidney, stomach and also in pheochromocytoma, melanoma, ductal carcinoma and carcinoid tumor.
[0042]Transfection of AIB1 into cells resulted in marked enhancement of estrogen-dependent transcription. These observations indicated that AIB1 functions as a co-activator of steroid hormone receptors such as ER (including estrogen receptor a (ERα) and estrogen receptor β (ERβ)), androgen receptor (e.g., expressed in prostate cells), retinoid receptor (e.g., isoforms α, γ, and retinoid X receptor (RXR)), progesterone receptor (e.g., expressed in breast cells), mineralocorticoid receptor (implicated in salt metabolism disorders), vitamin D receptor (implicated in calcium metabolism disorders), thyroid hormone receptor (e.g, thyroid hormone receptor α), or glucocorticoid receptor (e.g., expressed in spleen and thymus cells). The altered expression of AIB1 contributes to the initiation and progression of steroid hormone-responsive cancers by increasing the transcriptional activity of the steroid receptor.
[0043]A substantially pure DNA which includes an AIB1-encoding polynucleotides (or the complement thereof) is claimed. By "substantially pure DNA" is meant DNA that is free of the genes which, in the naturally-occurring genome of the organism from which the DNA of the invention is derived, flank the AIB1 gene. The term therefore includes, for example, a recombinant DNA which is incorporated into a vector, into an autonomously replicating plasmid or virus, or into the genomic DNA of a prokaryote or eukaryote at a site other than its natural site; or which exists as a separate molecule (e.g., a cDNA or a genomic or cDNA fragment produced by PCR or restriction endonuclease digestion) independent of other sequences. It also includes a recombinant DNA which is part of a hybrid gene encoding an additional polypeptide sequence. Preferably, the polypeptide includes a Per/Arnt/Sim (PAS) domain (LLQALDGFLFVVNRDGNIVFVSENVTQYLQYKQEDLVNTSVYNILHEEDRKDFLKNLPKSTVN GVSWTNETQRQKSHTFNCRMLMKTPHDILEDINASPEMRQRYETMQCFALSQPRAMMEEGED LQSCMICVARRITTGERTFPSNPESFITRHDLSGKVVNIDTNSLRSSMRPGFEDIIRRCIQ; SEQ. I.D. NO. 2) and/or a basic helix-loop-helix
(bHLH) domain (RKRKLPCDTPGQGLTCSGEKRRREQESKYIEELAELISANLSDIDNFNVKPDKCA ILKETVRQIRQIKEQGKT; SEQ. I.D. NO. 3); more preferably, the AIB1 polypeptide includes the amino acid sequence of the entire naturally-occurring AIB1 protein (FIG. 1; SEQ. I.D. NO. 4). Preferably, the peptide includes an ER-interacting domain of AIB1 (e.g., a domain comprising approximately amino acids 300 to 1250:
CIQRFFSLNDGQSWSQKRHYQEAYLNGHAETPVYRFSLADGTIVTAQTKSKLF
RNPVTNDRHGFVSTHFLQREQNGYRPNPNPVGQGIRPPMAGCNSSVGGMSMS
[0044]PNQGLQMPSSRAYGLADPSTTGQMSGARYGGSSNIASLTPGPGMQSPSSYQNNNYGLNMSSPPH GSPGLAPNQQNIMISPRNRGSPKIASHQFSPVAGVHSPMASSGNTGNHSFSSSSLSALQAISEGVGT SLLSTLSSPGPKLDNSPNMNITQPSKVSNQDSKSPLGFYCDQNPVESSMCQSNSRDHLSDKESKES SVEGAENQRGPLESKGHKKLLQLLTCSSDDRGHSSLTNSPLDSSCKESSVSVTSPSGVSSSTSGGV SSTSNMHGSLLQEKHRILHKLLQNGNSPAEVAKITAEATGKDTSSITSCGDGNVVKQEQLSPKKK ENNALLRYLLDRDDPSDALSKELQPQVEGVDNKMSQCTSSTIPSSSQEKDPKIKTETSEEGSGDLD NLDAILGDLTSSDFYNNSISSNGSHLGTKQQVFQGTNSLGLKSSQSVQSIRPPYNRAVSLDSPVSV GSSPPVKNISAFPMLPKQPMLGGNPRMMDSQENYGSSMGGPNRNVTVTQTPSSGDWGLPNSKA GRMEPMNSNSMGRPGGDYNTSLPRPALGGSIPTLPLRSNSIPGARPVLQQQQQMLQMRPGEIPM GMGANPYGQAAASNQLGSWPDGMLSMEQVSHGTQNRPLLRNSLDDLVGPPSNLEGQSDERAL LDQLHTLLSNTDATGLEEIDRALGIPELVNQGQALEPKQDAFQGQEAAVMMDQKAGLYGQTYP AQGPPMQGGFHLQGQSPSFNSMMNQMNQQGNFPLQGMHPRANIMRPRTNTPKQLRMQLQQR LQGQQFLNQSRQALELKMENPTAGGAAVMRPMMQPQQGFLNAQMVAQRSRELLSHHFRQQR VAMMMQQQQQQQ (SEQ. I.D. NO. 8). A cell containing substantially purified AIB1-encoding DNA is also within the invention.
[0045]The invention also includes a substantially pure DNA which contains a polynucleotide which hybridizes at high stringency to an AIB1 cDNA having the sequence of SEQ. I.D. NO. 1, or the complement thereof and a substantially pure DNA which contains a nucleotide sequence having at least 50% (for example at least 75%, 90%, 95%, or 98-100%) sequence identity to SEQ. I.D. NO. 1, provided the nucleotide sequence encodes a polypeptide having the biological activity of a AIB1 polypeptide. By "biological activity" is meant steroid receptor co-activator activity. For example, allelic variations of the naturally-occurring AIB1-encoding sequence (SEQ. I.D. NO. 1) are encompassed by the invention. Sequence identity can be determined by comparing the nucleotide sequences of two nucleic acids using the BLAST sequence analysis software, for instance, the NCBI gapped BLAST 2.0 program set to default parameters. This software is available from The National Center for Biotechnology Information (www.ncbi.nlm,nih.gov/BLAST).
[0046]Hybridization is carried out using standard techniques such as those described in Ausubel et al., Current Protocols in Molecular Biology, John Wiley & Sons, (1989). "High stringency" refers to DNA hybridization and wash conditions characterized by high temperature and low salt concentration, e.g., wash conditions of 65° C. at a salt concentration of approximately 0.1×SSC. "Low" to "moderate" stringency refers to DNA hybridization and wash conditions characterized by low temperature and high salt concentration, e.g. wash conditions of less than 60° C. at a salt concentration of at least 1.0×SSC. For example, high stringency conditions may include hybridization at about 42° C., and about 50% formamide; a first wash at about 65° C., about 2×SSC, and 1% SDS; followed by a second wash at about 65° C. and about 0.1%×SSC. Lower stringency conditions suitable for detecting DNA sequences having about 50% sequence identity to an AIB1 gene are detected by, for example, hybridization at about 42° C. in the absence of formamide; a first wash at about 42° C., about 6×SSC, and about 1% SDS; and a second wash at about 50° C., about 6×SSC, and about 1% SDS.
[0047]A substantially pure DNA including (a) the sequence of SEQ ID NO. 1 or (b) a degenerate variant thereof is also within the invention. The AIB1-encoding DNA is preferably operably linked to regulatory sequences (including, e.g., a promoter) for expression of the polypeptide.
[0048]By "operably linked" is meant that a coding sequence and a regulatory sequence(s) are connected in such a way as to permit gene expression when the appropriate molecules (e.g., transcriptional activator proteins) are bound to the regulatory sequence(s).
[0049]The invention also includes a substantially pure human AIB1 polypeptide or fragment thereof. The AIB1 fragment may include an ER-interaction domain such as one having the amino acid sequence of SEQ. I.D. NO. 8. Alternatively, the fragment may contain the amino acid sequence of SEQ. I.D. NOS. 2, 3, or 4.
[0050]Screening methods to identify candidate compounds which inhibit estrogen-dependent transcription, AIB1 expression, or an AIB1/ER interaction (and as a result, proliferation of steroid hormone-responsive cancer cells) are within the scope of the invention. For example, a method of identifying a candidate compound which inhibits ER-dependent transcription is carried out by contacting the compound with an AIB1 polypeptide and determining whether the compound binds to the polypeptide. Binding of the compound to the polypeptide indicates that the compound inhibits ER-dependent transcription, and in turn, proliferation of steroid hormone-responsive cancer cells. Preferably, the AIB1 polypeptide contains a PAS domain or a bHLH domain. Alternatively, the method is carried out by contacting the compound with an AIB1 polypeptide and an ER polypeptide and determining the ability of the compound to interfere with the binding of the ER polypeptide with the AIB1 polypeptide. A compound which interferes with an AIB1/ER interaction inhibits ER-dependent transcription.
[0051]A method of screening a candidate compound which inhibits an interaction of an AIB1 polypeptide with an ER polypeptide in a cell includes the steps of (a) providing a GAL4 binding site linked to a reporter gene; (b) providing a GAL4 binding domain linked to either (i) an AIB1 polypeptide or (ii) an ER polypeptide; (c) providing a GAL4 transactivation domain II linked to the ER polypeptide if the GAL4 binding domain is linked to the AIB1 polypeptide or linked to the AIB1 polypeptide if the GAL4 binding domain is linked to the ER polypeptide; (d) contacting the cell with the compound; and (e) monitoring expression of the reporter gene. A decrease in expression in the presence of the compound compared to that in the absence of the compound indicates that the compound inhibits an interaction of an AIB1 polypeptide with the ER polypeptide.
[0052]Diagnostic methods to identify an aberrantly proliferating cell, e.g., a steroid hormone-responsive cancer cell such as a breast cancer cell, ovarian cancer cell, or prostate cancer cell, are also included in the invention. For example, a method of detecting an aberrantly proliferating cell in a tissue sample is carried out by determining the level of AIB1 gene expression in the sample. An increase in the level of gene expression compared to that in a normal control tissue indicates the presence of an aberrantly proliferating cell. AIB1 gene expression is measured using an AIB1 gene-specific polynucleotides probe, e.g. in a Northern assay or polymerase chain reaction (PCR)-based assay, to detect AIB1 mRNA transcripts. AIB1 gene expression can also be measured using an antibody specific for an AIB1 gene product, e.g., by immunohistochemistry or Western blotting.
[0053]Aberrantly proliferating cells, e.g., cancer cells, in a tissue sample may be detected by determining the number of cellular copies of an AIB1 gene in the tissue. An increase in the number of gene copies in a cell of a patient-derived tissue, compared to that in normal control tissue indicates the presence of a cancer. A copy number greater than 2 (the normal diploid copy number) is indicative of an aberrantly proliferative cell. Preferably, the copy number is greater than 5 copies per diploid genome, more preferably 10 copies, more preferably greater than 20, and most preferably greater than 25 copies. An increase in copy number compared to the normal diploid copy number indicates that the tissue sample contains aberrantly proliferating steroid hormone-responsive cancer cells. AIB1 copy number is measured by fluorescent in situ hybridization (FISH), Southern hybridization techniques, and other methods well known in the art (Kallioniemi et al., PNAS 91: 2156-2160 (1994); Guan et al., Nature Genetics 8: 155-161 (1994); Tanner et al., Clin. Cancer Res. 1: 1455-1461 (1995); Guan et al., Cancer Res. 56: 3446-3450 (August 1996); Anzick et al., Science 277: 965-968 (August 1997)).
[0054]Aberrantly proliferating cells can also be identified by genetic polymorphisms in the polyglutamine tract of AIB1, e.g., variations in the size of this domain which alter AIB1 co-activator activity.
[0055]The invention also includes methods of treating a mammal, e.g., a human patient. For example, a method of reducing proliferation of a steroid hormone-responsive cancer cell, e.g., an estrogen-responsive breast cancer cell, in a mammal is carried out by administering to the mammal a compound which inhibits expression of AIB1. The compound reduces transcription of AIB1-encoding DNA in the cell. Alternatively, the compound reduces translation of an AIB1 mRNA into an AIB1 gene product in the cell. For example, translation of AIB1 mRNA into an AIB1 gene product is inhibited by contacting the mRNA with antisense polynucleotides complementary to the AIB1 mRNA.
[0056]A method of inhibiting ER-dependent transcription in a breast cell of a mammal is carried out by administering an effective amount of an AIB1 polypeptide or a peptide mimetic thereof to the mammal. Preferably, the polypeptide inhibits an AIB1/ER interaction; more preferably, the polypeptide contains an ER-interacting domain; a PAS domain or a bHLH domain of AIB1. By binding to ER, such a polypeptide inhibits binding of AIB1 to ER, thereby inhibiting ER-dependent transcription.
[0057]The invention also includes antibodies, e.g., a monoclonal antibody or polyclonal antisera, which bind specifically to AIB1. The term "antibody" as used in this invention includes whole antibodies as well as fragments thereof, such as Fab, Fab', F(ab')2, and Fv which bind to an AIB1 epitope. These antibody fragments are defined as follows: (1) Fab, the fragment which contains a monovalent antigen-binding fragment of an antibody molecule produced by digestion of whole antibody with the enzyme papain to yield an intact light chain and a portion of one heavy chain; (2) Fab', the fragment of an antibody molecule obtained by treating whole antibody with pepsin, followed by reduction, to yield an intact light chain and a portion of the heavy chain; two Fab' fragments are obtained per antibody molecule; (3) (Fab')2, the fragment of the antibody obtained by treating whole antibody with the enzyme pepsin without subsequent reduction; F(ab')2, a dimer of two Fab' fragments held together by two disulfide bonds; (4) Fv, a genetically engineered fragment containing the variable region of the light chain and the variable region of the heavy chain expressed as two chains; and (5) single chain antibody ("SCA"), a genetically engineered molecule containing the variable region of the light chain, the variable region of the heavy chain, linked by a suitable polypeptide linker as a genetically fused single chain molecule. Methods of making these fragments are routine.
[0058]Also within the invention is a method of identifying a tamoxifen-sensitive patient (one who is likely to respond to tamoxifen treatment by a reduction in rate of tumor growth) wherein the method includes the steps of (a) contacting a patient-derived tissue sample with tamoxifen; and (b) determining the level of AIB1 gene expression or amplification in the sample. An increase in the level of expression or gene copy number compared to the level or cellular copy number in normal control tissue indicates that the patient is tamoxifen-sensitive.
[0059]AIB1 gene expression is measured using an AIB1 gene-specific polynucleotide probe, e.g., in a Northern blot or PCR-based assay to detect AIB1 mRNA transcripts or in a Southern blot or FISH assay to detect amplification of the gene (which correlates directly with AIB1 gene expression). Alternatively, AIB1 gene expression is measured by detecting an AIB1 gene product, e.g., using an AIB1-specific antibody.
[0060]Transgenic mammals, e.g., mice, which overexpress an AIB1 gene product, e.g., by virtue of harboring multiple copies of AIB1-encoding DNA, are also within the invention.
[0061]"Transgenic" as used herein means a mammal which bears a transgene, a DNA sequence which is inserted by artifice into an embryo, and which then becomes part of the genome of the mammal that develops from that embryo. Any non-human mammal which may be produced by transgenic technology is included in the invention; preferred mammals include, mice, rats, cows, pigs, sheep, goats, rabbits, guinea pigs, hamsters, and horses.
[0062]By "transgene" is meant DNA which is partly or entirely heterologous (i.e., foreign) to the transgenic mammal, or DNA homologous to an endogenous gene of the transgenic mammal, but which is inserted into the mammal's genome at a location which differs from that of the natural gene.
[0063]Also within the invention is a knockout mutant, for instance a knockout mouse wherein the mouse has had at least one copy of the AIB1 gene (also called the pCIP gene in mice) deleted from its genome. Such a knockout mutant would be useful in research, for instance the phenotype gives insight into the physiological role of AIB1. Complementation experiments using such a knockout mutant can be used to identify other genes and proteins that make up for the lack of AIB1 in the mutant to restore wild-type phenotype.
[0064]Also within the invention is a mutant, such as a mouse, which contains more than the normal number of copies of the AIB1 (pCIP) gene, either integrated into a chromosome, for instance as a pro-virus, or in an extra-chromosomal element, such as on a plasmid.
[0065]Also within the invention is a mutant, for example, a mouse, which contains the AIB1 (pCIP) gene driven by a non-native promoter, such as a constitutive or an inducible promoter, such as the mouse mammary tumor virus (MMTV) promoter.
[0066]The invention also includes methods of treatment for cancers the growth of which involves alternations of signaling pathways involving p300 and/or CBP. For example, AIB1 (pCIP) may be contacted with a molecule that binds to AIB1 and inhibits AIB1's interaction with p300, thereby disrupting signaling of this pathway and reducing transcription of molecules whose transcription is positively regulated by this pathway; thereby reducing tumor growth.
EXAMPLE 1
Cloning and Expression of AIB1
A. Cloning of AIB1
[0067]Chromosome microdissection and hybrid selection techniques were used to isolate probes and clone gene sequences which map to chromosome 20q, one of the recurrent sites of DNA amplification in breast cancer cells identified by molecular cytogenetics (Kallioniemi et al., PNAS 91: 2156-2160 (1994); Guan et al., Nature Genetics 8: 155-161 (1994); Tanner et al., Clin. Cancer Res. 1: 1455-1461 (1995); Guan et al., Cancer Res. 56: 3446-3450 (August 1996); Anzick et al., Science 277: 965-968 (August 1997)). AIB1 is a member of the SRC-1 family of nuclear receptor (NR) co-activators. AIB1 functions to enhance ER-dependent transcription. SRC-1 and the closely related TIF2 are steroid receptor co-activators with an affinity for NRs. The mouse ortholog of human AIB1 is called pCIP. In this application pCIP and AIB1 will be used synonymously unless the contrary is clearly expressed.
[0068]To characterize AIB1, the full length cDNA was cloned and sequenced. An AIB1 specific primer N8F1 (5'-TCATCACTTCCGACAACAGAGG-3'; SEQ. I.D. NO. 5) was biotinylated and used to capture cDNA clones from a human lung cDNA library (Gibco, BRL) using the GENETRAPPER cDNA Positive Selection System (Gibco, BRL). The largest clone (5.8 kb), designated pCMVSPORT-B11, was selected for sequence analysis. To obtain full-length AIB1-encoding DNA, a random-primed library from BT474 was constructed in bacteriophage λ-Zap (Stratagene) and hybridized with a 372 bp 32P-labeled PCR product amplified from a human spleen cDNA library using primers designed form the 5' sequence of pCMVSPORT-B11, PM-U2 (5'-CCAGAAACGTCACTATCAAG-3', forward primer; SEQ. I.D. NO. 6) and B11-11RA (5'-TTACTGGAACCCCCATACC-3', reverse primer; SEQ. I.D. NO. 7). Plasmid rescue of 19 positive clones yielded a clone, pBluescript-R22, which overlapped pCMVSPORT-B11 and contained the 5' end of the coding region. To generate a full length AIB1 clone, the 4.85 kb HindIII/XhoI fragment of pCMVSPORT-B11 was subcloned into HindIII/XhoI sites of pBluescript-R22. The 4.84 kb NotI/NheI fragment of the full length clone containing the entire coding region was then subcloned into the NotI/XbaI sites of the expression vector, pcDNA3.1 (Invitrogen), generating pcDNA3.1-AIB1.
[0069]The cloned DNA sequence (SEQ. I.D. No. 1) revealed an open reading frame (beginning at the underlined "ATG") encoding a protein of 1420 amino acids with a predicted molecular weight of 155 kDa (FIG. 1A). Database searches with BLASTP identified a similarity of AIB1 with TIF2 (45% protein identity) and SRC-1 (33% protein identity). Like TIF2 and SRC-1, AIB1 contains a bHLH domain preceding a PAS domain, serine/threonine-rich regions, and a charged cluster (FIG. 1B). There is also a glutamine-rich region which, unlike SRC-1 and TIF2, contains a polyglutamine tract (FIG. 1B). The polyglutamine tract of AIB1 is subject to genetic polymorphism. Variations in the size of this domain alter AIB1 co-activator activity.
B. Expression of AIB1
[0070]Amplification and expression of AIB1 in several ER positive and negative breast and ovarian cancer cell lines was examined. Established breast cancer cell lines used in the experiments described below (see, e.g., FIG. 2) were obtained from the American Type Culture Collection (ATCC): BT-474, MCF-7, T47D, MDA-MB-361, MDA-MB-468, BT-20, MDA-MB-436, and MDA-MB-453; the Arizona Cancer Center (ACC): UACC-812; or the National Cancer Institute (NCI): ZR75-1.
[0071]AIB1 gene copy number was determined by FISH. For FISH analysis, interphase nuclei were fixed in methanol:acetic acid (3:1) and dropped onto microscope slides. AIB1 amplification was detected in the breast cancer cell line ZR75-1, the ovarian cancer cell line BG-1, and two uncultured breast cancer samples. Intra-chromosomal amplification of AIB1 was apparent in metaphase chromosomes of ZR75-1 and BG1. Numerous copies of AIB1 were resolved in the adjacent interphase nuclei. Extrachromosomal copies (e.g., in episomes or double minute chromosomes) of AIB1 have also been detected. The Spectrum-Orange (Vysis) labeled AIB1 P1 probe was hybridized with a biotinylated reference probe for 20q11 (RMC20P037) or a fluorescein labeled probe for 20p (RMC20C039).
[0072]High level amplification of AIB1 (greater than 20 fold), similar to that observed in BT-474 and MCF-7, was seen in two additional ER-positive cell lines, breast carcinoma ZR75-1, and ovarian carcinoma BG-1 (see FIG. 2). Interphase FISH studies demonstrated that amplification of chromosome 20q in breast cancer is complex, involving several distinct variably co-amplified chromosomal segments derived from 20q11, 20q12, and 20q13. Probes for the 20q11 and 20q13 regions of amplification did not detect amplification in ZR75-1 and BG-1, suggesting that amplification of AIB1 (which maps to 20q12) occurred independently in these cell lines.
[0073]To determine if AIB1 amplification also occurred in uncultured cells from patient biopsies, breast cancer specimens were screened for AIB1 amplification by interphase FISH. In two of 16 specimens analyzed, high AIB1 copy number (up to 25 copies/cell) was detected. Both tumor specimens tested came from post-menopausal patients and were ER/PR positive. One of the specimens was obtained from a metastatic tumor of a patient who subsequently responded favorably to tamoxifen treatment.
[0074]AIB1 expression was also examined in cells with and without AIB1 amplification and compared to expression of ER, SRC-1 and TIF2 by Northern blotting. In accordance with its amplification status, AIB1 was highly overexpressed in BT-474, MCF-7, ZR75-1, and BG-1 (FIG. 2). Three of the four cell lines exhibiting AIB1 overexpression also demonstrated prominent ER expression, while two others displayed lower but detectable ER expression (BT-474 and BT-20). FIG. 2 also shows that the expression of TIF2 and SRC-1 remained relatively constant in all cell lines tested. Taken together, these observations demonstrate that AIB1 amplification is associated with significant overexpression of AIB1 gene product. The correlation of elevated AIB1 expression with ER positivity in tumors indicates that AIB1 is a component of the estrogen signaling pathway, the amplification of which is selected during cancer development and progression.
[0075]To determine whether expression of AIB1 increases ER ligand-dependent transactivation, transient transfection assays were performed. The effect of increasing levels of AIB1 on transcription of an ER dependent reporter was measured. The results demonstrated that co-transfection of AIB1 led to a dose dependent increase in estrogen-dependent transcription (FIG. 3). This effect was not observed when the estrogen antagonist, 4-hydroxytamoxifen (4-OHT), was substituted for 17β-estradiol or when the estrogen response element (ERE) was removed from the reporter plasmid (FIG. 3). A modest increase in basal transcription levels was observed with higher concentrations of AIB1 even in the absence of an ERE suggesting that AIB1 may have an intrinsic transactivation function. These results demonstrate that, like the closely related TIF2 and SRC-1, AIB1 functions as an ER co-activator.
EXAMPLE 2
Characterization of AIB1
A. Functional Domains of AIB1
[0076]TIF-2, SRC-1, and AIB1 are characterized by highly conserved N-terminal bHLH and PAS domains. The PAS region functions as a protein dimerization interface in the mammalian aryl hydrocarbon receptor and the aryl hydrocarbon receptor nuclear transporter proteins, as well as the Drosophila transcription factors sim and per. The PAS region (SEQ. I.D. NO. 2) of AIB1 functions as a protein interaction domain, mediating binding between AIB1 and other proteins. However, steroid hormone activators lacking the PAS domain are capable of interacting with nuclear steroid hormone receptors. The highly conserved bHLH domain (SEQ. I.D. NO. 3) participates in protein interactions which mediate or modulate transmission of the hormone signal to the transcriptional apparatus. The ER-interacting domain (SEQ. I.D. NO. 8) mediates binding of AIB1 with a steroid hormone receptor protein.
[0077]AIB1 also interacts with the transcriptional integrators CREB binding protein (CBP) and p300. These transcriptional integrators interact directly with the basal transcriptional machinery. The CBP/p300 receptor association domain of AIB1 does not encompass the bHLH/PAS regions.
B. Purification of Gene Products
[0078]DNA containing a sequence that encodes part or all of the amino acid sequence of AIB1 can be subcloned into an expression vector, using a variety of methods known in the art. The recombinant protein can then be purified using standard methods. For example, a recombinant polypeptide can be expressed as a fusion protein in procaryotic cells such as E. coli. Using the maltose binding protein fusion and purification system (New England Biolabs), the cloned human cDNA sequence is inserted downstream and in frame of the gene encoding maltose binding protein (malE). The malE fusion protein is overexpressed in E. coli and can be readily purified in quantity. In the absence of convenient restriction sites in the human cDNA sequence, PCR can be used to introduce restriction sites compatible with the pMalE vector at the 5' and 3' end of the cDNA fragment to facilitate insertion of the cDNA fragment into the vector. Following expression of the fusion protein, it can be purified by affinity chromatography. For example, the fusion protein can be purified by virtue of the ability of the maltose binding protein portion of the fusion protein to bind to amylase immobilized on a column.
[0079]To facilitate protein purification, the pMalE plasmid contains a factor Xa cleavage site upstream of the site into which the cDNA is inserted into the vector. Thus, the fusion protein purified as described above can be cleaved with factor Xa to separate the maltose binding protein portion of the fusion protein from recombinant human cDNA gene product. The cleavage products can be subjected to further chromatography to purify recombinant polypeptide from the maltose binding protein. Alternatively, an antibody specific for the desired recombinant gene product can be used to purify the fusion protein and/or the gene product cleaved from the fusion protein. Many comparable commercially available fusion protein expression systems can be utilized similarly.
[0080]AIB1 polypeptides can also be expressed in eucaryotic cells, e.g., yeast cells, either alone or as a fusion protein. For example, a fusion protein containing the GAL4 DNA-binding domain or activation domain fused to a functional domain of AIB1, e.g., the PAS domain, the bHLH domain, or the ER-interacting domain, can be expressed in yeast cells using standard methods such as the yeast two hybrid system described below. Alternatively, AIB1 polypeptides can be expressed in COS-1 cells using methods well known in the art, e.g., by transfecting a DNA encoding an AIB1 polypeptide into COS-1 cells using, e.g., the Lipofectamine transfection protocol described below, and culturing the cells under conditions suitable for protein expression.
EXAMPLE 3
Detection of AIB1
A. Detection of Nucleotides Encoding AIB1
[0081]Determination of gene copy number in cells of a patient-derived sample is known in the art. For example, AIB1 amplification in cancer-derived cell lines as well as uncultured breast cancer cells was carried out using bicolor FISH analysis as follows. A genomic P1 clone containing AIB1 was labeled with Spectrum Orange-dUTP (Vysis) using the BioPrime DNA Labeling System (Gibco BRL). A 20q11 P1 clone was labeled with Biotin-16-dUTP (BMB) using nick translation. Fluorescent images were captured using a Zeiss axiophot microscope equipped with a CCD camera and IP Lab Spectrum software (Signal Analytics). Interphase FISH analysis of uncultured breast cancer samples was performed using known methods (Kallioniemi et al., PNAS 91: 2156-2160 (1994); Guan et al., Nature Genetics 8: 155-161 (1994); Tanner et al., Clin. Cancer Res. 1: 1455-1461 (1995); Guan et al., Cancer Res. 56: 3446-3450 (August 1996); Anzick et al., Science 277: 965-968 (August 1997)). Alternatively, standard Southern hybridization techniques can be employed to evaluate gene amplification. For example, Southern analysis is carried out using a non-repetitive fragment of genomic AIB1 DNA, e.g., derived from the 20q11 P1 clone described above or another AIB1 gene-containing genomic clone, as a probe.
[0082]The level of gene expression may be measured using methods known in the art, e.g., in situ hybridization, Northern blot analysis, or Western blot analysis using AIB1-specific monoclonal or polyclonal antibodies. AIB1 gene transcription was measured using Northern analysis. For example, the data shown in FIG. 2 was obtained as follows. The blot was hybridized sequentially with a probe (ER, AIB1, TIF2, SRC-1, or β-actin as indicated to the left of the photograph). AIB1 expression was compared to that of ER, TIF2, and SRC-1. cDNA clones were obtained from Research Genetics [TIF2 (clone 132364, GenBank accession no. R25318); SRC-1 (clone 418064, GenBank accession no. W90426)], the American Type Culture Collection (pHEGO-hyg, ATCC number 79995), or Clontech (β actin). The AIB1 probe was a 2.2 kb NotI/SacI fragment of pCMVSPORT-B11. The β-actin probe was used as a control for loading error. To avoid cross-hybridization between these related genes and to match signal intensities, similar sized probes from the 3'UTRs of AIB1, TIF2, and SRC-1 were utilized. Each of these probes detected a signal in normal mammary RNA on longer exposure. Electrophoresis, transfer and hybridization of 15 μg total RNA was performed by standard methods.
B. Detection of AIB1 Gene Products
[0083]AIB1 polypeptides to be used as antigens to raise AIB1-specific antibodies can be generated by methods known in the art, e.g., proteolytic cleavage, de novo synthesis, or expression of a recombinant polypeptide from the cloned AIB1 gene or a fragment thereof. AIB1-specific antibodies are then produced using standard methodologies for raising polyclonal antisera and making monoclonal antibody-producing hybridoma cell lines (see Coligan et al., eds., Current Protocols in Immunology, 1992, Greene Publishing Associates and Wiley-Interscience). To generate monoclonal antibodies, a mouse is immunized with an AIB1 polypeptide, antibody-secreting B cells isolated from the mouse, and the B cells immortalized with a non-secretory myeloma cell fusion partner. Hybridomas are then screened for production of an AIB1-specific antibody and cloned to obtain a homogenous cell population which produces a monoclonal antibody.
[0084]For administration to human patients, antibodies, e.g., AIB1 specific monoclonal antibodies, can be humanized by methods known in the art. Antibodies with a desired binding specificity can be commercially humanized (Scotgene, Scotland; Oxford Molecular, Palo Alto, Calif.).
EXAMPLE 4
Detection of AIB1-Related Cell Proliferative Disorders
A. Diagnostic and Prognostic Methods
[0085]The invention includes a method of detecting an aberrantly proliferating cell, e.g., a steroid hormone-responsive cancer cell such as a breast cancer cell, an ovarian cancer cell, colon cancer cell, or prostate cancer cell, by detecting the number of AIB1 gene copies in the cell and/or the level of expression of the AIB1 gene product. AIB1 gene amplification or gene expression in a patient-derived tissue sample is measured as described above and compared to the level of amplification or gene expression in normal non-cancerous cells. An increase in the level of amplification or gene expression detected in the patient-derived biopsy sample compared to the normal control is diagnostic of a diseased state, i.e., the presence of a steroid hormone responsive cancer.
[0086]Because of the importance of estrogen exposure to mammary carcinogenesis and of anti-estrogen treatment in breast cancer therapy, such assays are also useful to determine the frequency of alterations of AIB1 expression in pre-malignant breast lesions (e.g. ductal carcinoma in situ) and during the progression from hormone dependent to hormone independent tumor growth.
[0087]The diagnostic methods of the invention are useful to determine the prognosis of a patient and estrogen responsive status of a steroid hormone-responsive cancer.
[0088]AIB1 expression can also be measured at the protein level by detecting an AIB1 gene products with an AIB1-specific monoclonal or polyclonal antibody preparation.
B. Diagnosis of Tamoxifen-Sensitivity
[0089]Overexpression of AIB1, e.g., as a result of AIB1 gene amplification, in steroid hormone-responsive cancers can predict whether the cancer is treatable with anti-endocrine compositions, e.g., tamoxifen. AIB1 amplification or overexpression in a patient-derived tissue sample compared to a normal (non-cancerous) tissue indicates tumor progression.
[0090]Absence of AIB1, e.g., loss of all or part of the AIB1 gene, but retention of ER-positivity in steroid hormone-responsive cancers predicts failure or poor responsiveness to anti-endocrine therapy, e.g., administration of anti-estrogen compositions such as tamoxifen. Since loss of AIB1 expression in a cancer cell may indicate a disruption of the ER signal transduction pathway, anti-estrogen therapy may be ineffective to treat such cancers. Patients identified in this manner (who would otherwise be treated with anti-estrogens) would be treated with alternative therapies.
[0091]Loss of estrogen receptor in recurrent breast caner is also associated with poor response to endocrine therapy. Up to 30% to 40% of metastases from hormone receptor-positive primary breast cancer do not respond to endocrine therapy. The frequency of hormone receptor status changes between primary and recurrent tumors and whether such a change might explain unresponsiveness to endocrine therapy was examined. Primary breast cancer samples and matched asynchronous recurrences were studied from 50 patients who had not received any adjuvant therapy. ER and progesterone receptor (PR) status was determined immunohistochemically from histologically representative formalin-fixed paraffin-embedded tumor samples. ER status was ascertained by mRNA in situ hybridization. Thirty-five (70%) of 50 primary tumors were positive for ER and 30 (60%) for PR. Hormone receptor status of the recurrent tumor differed from that of the primary tumor in 18 cases (36%). Discordant cases were due to the loss of ER (n=6), loss of PR (n=6), or loss of both receptors (n=6). Receptor-negative primary tumors were always accompanied by receptor-negative recurrences. Among 27 patients with ER-positive primary tumors, loss of ER was a significant predictor (P=0.0085) of poor response to subsequent endocrine therapy. Only one of eight patients (12.5%) with lost ER expression responded to tamoxifen therapy, whereas the response rate was 74% (14 of 19) for patients whose recurrent tumors retained ER expression. Loss of ER expression in recurrent breast cancer predicts poor response to endocrine therapy in primarily ER-positive patients. Evaluation of ER expression and/or AIB1 expression (or gene copy number) is useful to determine the most effective approach to treatment of steroid-responsive cancers.
EXAMPLE 5
Screening of Candidate Compounds
A. In Vitro Assays
[0092]The invention includes methods of screening to identify compounds which inhibit the interaction of AIB1 with ER, thereby decreasing estrogen dependent transcription which leads to aberrant cell proliferation. A transcription assay is carried out in the presence and absence of the candidate compound. A decrease in transcription in the presence of the compound compared to that in its absence indicates that the compound blocks an AIB1/ER interaction and inhibits estrogen dependent transcription.
[0093]To determine the effect of AIB1 on estrogen-dependent transcription, an ER reporter plasmid can be used. The transcription assays described herein were conducted as follows. COS-1 cells were grown and maintained in phenol-red free DMEM medium supplemented with 10% charcoal-stripped fetal bovine serum. Cells were plated into 6-well culture dishes at 1.5×105 cells/well and allowed to grow overnight. Transfection of cells with the ER reporter plasmid was performed with Lipofectamine (Gibco, BRL) following the manufacturer's protocol. Three ng pRL-CMV were used as an internal control for transfection efficiency. Ligand or ethanol vehicle was added 234 hours post-transfection and cell lysates were harvested 48 hours post-transfection. Reporter activities were determined using the Dual-Luciferase Reporter Assay System (Promega) and the results expressed in relative luminescence units (RLU; luciferase/Renilla luciferase). pRL-CMV and pGL3-promoter were obtained from Promega. pHEGO-hyg was obtained from ATCC. The ER reporter pGL3.luc.3ERE contains three tandem copies of the ERE upstream from the SV40 promoter driving the luciferase gene. Standard mammalian expression vectors were utilized. Empty pcDNA3 vector was added to each of the pcDNA3.1-AIB1 dilutions to maintain constant amounts of plasmid DNA.
[0094]Compounds which inhibit the interaction of AIB1 with ER are also identified using a standard co-precipitation assay. AIB1/ER co-precipitation assays are carried out as follows. An AIB1 polypeptide and an ER polypeptide are incubated together to allow complex formation. One of the polypeptides is typically a fusion protein, e.g., GST-AIB1, and the other is tagged with a detectable label, e.g., 32P-labeled ER). After incubation, the complex is precipitated, e.g., using glutathione-Sepharose beads. The beads are washed, filtered through a glass fiber filter, and collected. The amount of co-precipitated 32P-label is measured. A reduction in the amount of co-precipitated label in the presence of a candidate compound compared to that in the absence of the candidate compound indicates that the compound inhibits an AIB1/ER interaction
[0095]Alternatively, a standard in vitro binding assay can be used. For example, one polypeptide, e.g., AIB1, can be bound to a solid support and contacted with the second polypeptide, e.g., ER. The amount of the second polypeptide which is retained on the solid support is then measured. A reduction in the amount of retained (second) polypeptide in the presence of a candidate compound compared to that in its absence indicates that the compound inhibits an AIB1/ER interaction. Techniques for column chromatography and coprecipitation of polypeptides are well known in the art.
[0096]An evaluation of AIB1/ER interaction and identification of compounds that blocks or reduces the interaction can also be carried out in vivo using a yeast two-hybrid expression system in which the activity of a transcriptional activator is reconstituted when the two proteins or polypeptides of interest closely interact or bind to one another.
[0097]The yeast GAL4 protein consists of functionally distinguishable domains. One domain is responsible for DNA-binding and the other for transcriptional activation. In the two-hybrid expression system, plasmids encoding two hybrid proteins, a first fusion protein containing the GAL4 DNA-binding domain fused to a first protein, e.g., AIB1, and the second fusion protein containing the GAL4 activation domain fused to a second protein, e.g., ER, are introduced into yeast. If the two proteins are able to interact with one another, the ability to activate transcription from promoters containing Gal4-binding sites upstream from an activating sequence from GALL (UASG) is reconstituted leading to the expression of a reporter gene. A reduction in the expression of the reporter gene in the presence of a candidate compound compared to that in the absence of the compound indicates that the compound reduces an AIB1/ER interaction.
[0098]A method of identifying a DNA-binding protein which regulates AIB1 transcription can be carried out as follows:
A DNA containing a cis-acting regulatory element can be immobilized on polymeric beads, such as agarose or acrylamide. A mixture of proteins, such as a cell lysate, is allowed to come in contact with and bind to the DNA. Following removal of non-binding proteins, specifically-bound proteins, are eluted with a competing DNA sequence which may be identical to the immobilized sequence. Specific binding of a protein to the DNA regulatory element indicates that the protein may regulate AIB1 transcription. Functional activity of the identified trans-acting factor can be confirmed with an appropriate functional assay, such as one which measures the level of transcription of a reporter gene having the cis-acting regulatory gene 5' to the transcription start site of AIB1.
[0099]A method of identifying a compound which decreases the level of AIB1 transcription can be accomplished by contacting an immobilized AIB1-derived cis-acting regulatory element with a trans-acting regulatory factor in the presence and absence of candidate compound. A detectable change, i.e., a reduction, in specific binding of the trans-acting factor to its DNA target indicates that the candidate compound inhibits AIB1 transcription.
[0100]In addition to interacting with ER, AIB1 also interacts with the transcriptional integrators CBP and p300. CBP and p300 participate in the basal transcriptional apparatus in a cell. Thus, another approach to inhibit signal transduction through AIB1 is to prevent the formation of or disrupt an interaction of AIB1 with CBP and/or p300. Compounds which inhibit signal transduction (and therefore cell proliferation) can be identified by contacting AIB1 (or a fragment thereof which interacts with CBP or p300) with CBP or p300 (or a fragment thereof containing an AIB1-interacting domain, e.g., a C-terminal fragment) in the presence and absence of a candidate compound. For example, a C-terminal fragment of CBP involved in steroid receptor co-activator interaction contains 105 amino acids in the Q-rich region of CBP (Kamei et al., 1996, Cell 85:403-414; Yao et al., 1996, Proc. Natl. Acad. Sci. USA 93:10626-10631; Hanstein et al., 1996, Proc. Natl. Acad. Sci. USA 93:11540-11545). A decrease in AIB1 interaction with CBP or p300 in the presence of a candidate compound compared to that its absence indicates that the compound inhibits AIB1 interaction with these transcriptional integrators, and as a result, AIB1-mediated signal transduction leading to DNA transcription and cell proliferation. Compounds which inhibit AIB1 interaction with transcriptional integrators can also be identified using a co-precipitation assay and the yeast two-hybrid expression system described above.
B. In Vivo Assays
[0101]Transgenic mice are made by standard methods, e.g., as described in Leder et al., U.S. Pat. No. 4,736,866, herein incorporated by reference, or Hogan et al., 1986 Manipulating the Mouse Embryo. Cold Spring Harbor Laboratory" New York.
[0102]Briefly, a vector containing a promoter operably linked to AIB1-encoding cDNA is injected into murine zygotes, e.g., C57BL/6J×DBA/2F2 zygotes. Incorporation of the transgene into murine genomic DNA is monitored using methods well known in the art of molecular biology, e.g., dot blotting tail DNA with a probe complimentary to the 3' region of the gene contained in the AIB1 transgene construct. Mice thus confirmed to harbor the transgene can then be used as founders. Animal lines are created by crossing founders with C57BL/6J mice (The Jackson Laboratory, Bar Harbor, Me.). AIB1 transgenic mice can be used to screen candidate compounds in vivo to identify compounds which inhibit aberrant cell proliferation, e.g., as measured by reduction tumor growth or metastasis. AIB1 transgenic mice are also useful to identify other genes involved in steroid hormone receptor-dependent cancers and to establish mouse cell lines which overexpress AIB1. AIB1-overexpressing cell lines are useful to screen for compounds that interfere with AIB1 function, e.g, by blocking the interaction of AIB1 with a ligand.
EXAMPLE 6
AIB1 Therapy
[0103]As discussed above, AIB1 is a novel member of the SRC-1 family of transcriptional co-activators. Amplification and overexpression of AIB1 in ER-positive breast and ovarian cancer cells and in breast cancer biopsies implicate this protein as a critical component of the estrogen response pathway. AIB1 overexpression results in increased ER-dependent transcriptional activity which confers a growth advantage of AIB1 amplification-bearing clones during the development and progression of estrogen-dependent cancers.
[0104]Compounds which inhibit or disrupt the interaction of an AIB1 gene product with a steroid hormone receptor, e.g., ER, are useful as anti-neoplastic agents for the treatment of patients suffering from steroid hormone-responsive cancers such as breast cancer, ovarian cancer, prostate cancer, and colon cancer. Likewise, compounds which disrupt interaction between AIB1 and p300 and/or CBP are also useful as anti-neoplastic agents.
[0105]AIB1 polypeptides or peptide mimetics of such polypeptides, e.g., those containing domains which interact with steroid hormone receptors, can be administered to patients to block the interaction of endogenous intracellular AIB1 and a steroid hormone receptor, e.g., ER in an aberrantly proliferating cell. A mimetic may be made by introducing conservative amino acid substitutions into the peptide. Certain amino acid substitutions are conservative since the old and the new amino acid share a similar hydrophobicity or hydrophylicity or are similarly acidic, basic or neutrally charged (Stryer "Biochemistry" 1975, Ch. 2, Freeman and Company, New York). Conservative substitutions replace one amino acid with another amino acid that is similar in size, hydrophobicity, etc. Examples of conservative substitutions are shown in the table below (Table 1).
TABLE-US-00001 TABLE 1 Original Residue Conservative Substitutions Ala ser Arg lys Asn gln, his Asp glu Cys ser Gln asn Glu asp Gly pro His asn; gln Ile leu, val Leu ile; val Lys arg; gln; glu Met leu; ile Phe met; leu; tyr Ser thr Thr ser Trp tyr Tyr trp; phe Val ile; leu
[0106]Variations in the cDNA sequence that result in amino acid changes, whether conservative or not, should be minimized in order to preserve the functional and immunologic identity of the encoded protein.
[0107]Compositions administered therapeutically include polypeptide mimetics in which one or more peptide bonds have been replaced with an alternative type of covalent bond which is not susceptible to cleavage by peptidases. Where proteolytic degradation of the peptides following injection into the subject is a problem, replacement of a particularly sensitive peptide bond with a noncleavable peptide mimetic yields a more stable and thus more useful therapeutic polypeptide. Such mimetics, and methods of incorporating them into polypeptides, are well known in the art. Similarly, the replacement of an L-amino acid residue with a D-amino acid residue is a standard way of rendering the polypeptide less sensitive to proteolysis. Also useful are amino-terminal blocking groups such as t-butyloxycarbonyl, acetyl, theyl, succinyl, methoxysuccinyl, suberyl, adipyl, azelayl, dansyl, benzyloxycarbonyl, fluorenylmethoxycarbonyl, methoxyazelayl, methoxyadipyl, methoxysuberyl, and 2,4,-dinitrophenyl.
[0108]AIB1 polypeptides or related peptide mimetics may be administered to a patient intravenously in a pharmaceutically acceptable carrier such as physiological saline. Standard methods for intracellular delivery of peptides can be used, e.g. packaged in liposomes. Such methods are well known to those of ordinary skill in the art. It is expected that an intravenous dosage of approximately 1 to 100 μmoles of the polypeptide of the invention would be administered per kg of body weight per day. The compositions of the invention are useful for parenteral administration, such as intravenous, subcutaneous, intramuscular, and intraperitoneal.
[0109]The therapeutic compositions of this invention may also be administered by the use of surgical implants which release the compounds of the invention. These devices could be readily implanted into the target tissue, e.g., a solid tumor mass, and could be mechanical or passive. Mechanical devices, such as pumps, are well known in the art, as are passive devices (e.g., consisting of a polymer matrix which contains therapeutic formulations; these polymers may slowly dissolve or degrade to release the compound, or may be porous and allow release via pores).
[0110]Antisense therapy in which a DNA sequence complementary to an AIB1 mRNA transcript is either produced in the cell or administered to the cell can be used to decrease AIB1 gene expression thereby inhibiting undesired cell proliferation, e.g., proliferation of steroid hormone-responsive cancer cells. An antisense polynucleotide, i.e., one which is complementary of the coding sequence of the AIB1 gene, is introduced into the cells in which the gene is overproduced. The antisense strand (either RNA or DNA) may be directly introduced into the cells in a form that is capable of binding to the transcripts. Alternatively, a vector containing a DNA sequence which, once within the target cells, is transcribed into the appropriate antisense mRNA, may be administered. An antisense nucleic acid which hybridizes to the coding strand of AIB1 DNA can decrease or inhibit production of an AIB1 gene product by associating with the normally single-stranded mRNA transcript, and thereby interfering with translation.
[0111]DNA is introduced into target cells of the patient with or without a vector or using standard vectors and/or gene delivery systems. Suitable gene delivery systems may include liposomes, receptor-mediated delivery systems, naked DNA, and viral vectors such as herpes viruses, retroviruses, and adenoviruses, among others. The DNA of the invention may be administered in a pharmaceutically acceptable carrier. Pharmaceutically acceptable carriers are biologically compatible vehicles which are suitable for administration to an animal e.g., physiological saline. A therapeutically effective amount is an amount of the nucleic acid of the invention which is capable of producing a medically desirable result in a patient. As is well known in the medical arts, dosage for any given patient depends upon many factors, including the patient's size, body surface area, age, the particular compound to be administered, sex, time and route of administration, general health, and other drugs being administered concurrently. Dosages will vary, but a preferred dosage for intravenous administration of a nucleic acid is from approximately 106 to 1022 copies of the nucleic acid molecule.
[0112]Determination of optimal dosage is well within the abilities of a pharmacologist of ordinary skill.
EXAMPLE 7
AIB1 Knockout and Overexpression Mouse Mutants
[0113]Mutants organism that underexpress or overexpress AIB1 are useful for research. Such mutants allow insight into the physiological and/or pathological role of AIB1 in a healthy and/or pathological organism. These mutants are said to be "genetically engineered," meaning that information in the form of nucleotides has been transferred into the mutant's genome at a location, or in a combination, in which it would not normally exist. Nucleotides transferred in this way are said to be "non-native." For example, a WAP promoter inserted upstream of a native AIB1 gene would be non-native. An extra copy of a mouse AIB1 gene present on a plasmid and transformed into a mouse cell would be non-native. Mutants may be, for example, produced from mammals, such as mice, that either overexpress AIB1 or underexpress AIB1 or that do not express AIB1 at all. Overexpression mutants are made by increasing the number of AIB1 genes in the organism, or by introducing an AIB1 gene into the organism under the control of a constitutive or inducible or viral promoter such as the mouse mammary tumor virus (MMTV) promoter or the whey acidic protein (WAP) promoter or the metallothionein promoter. Mutants that underexpress AIB1 may be made by using an inducible or repressible promoter, or by deleting the AIB1 gene, or by destroying or limiting the function of the AIB1 gene, for instance by disrupting the gene by transposon insertion.
[0114]Anti-sense genes may be engineered into the organism, under a constitutive or inducible promoter, to decrease or prevent AIB1 expression. A gene is said to be "functionally deleted" when genetic engineering has been used to negate or reduce gene expression to negligible levels. When a mutant is referred to in this application as having the AIB1 gene altered or functionally deleted, this reference refers to the AIB1 gene and to any ortholog of this gene, for instance "a transgenic animal wherein at least one AIB1 gene has been functionally deleted" would encompass the mouse ortholog of the AIB1 gene, pCIP. When a mutant is referred to as having "more than the normal copy number" of a gene, this means that it has more than the usual number of genes found in the wild-type organism, eg: in the diploid mouse or human.
[0115]A mutant mouse overexpressing AIB1 may be made by constructing a plasmid having the AIB1 gene driven by a promoter, such as the mouse mammary tumor virus (MMTV) promoter or the whey acidic protein (WAP) promoter. This plasmid may be introduced into mouse oocytes by microinjection. The oocytes are implanted into pseudopregnant females, and the litters are assayed for insertion of the transgene. Multiple strains containing the transgene are then available for study.
[0116]WAP is quite specific for mammary gland expression during lactation, and MMTV is expressed in a variety of tissues including mammary gland, salivary gland and lymphoid tissues. Many other promoters might be used to achieve various patterns of expression, e.g., the metallothionein promoter.
[0117]An inducible system may be created in which AIB1 is driven by a promoter regulated by an agent which can be fed to the mouse such as tetracycline. Such techniques are well known in the art.
[0118]A mutant knockout mouse from which the AIB1 (also called pCIP) gene is deleted was made by removing coding regions of the AIB1 gene from mouse embryonic stem cells. FIG. 5 shows the intron/exon structure for pCIP. Using this table, mutations can be targeted to coding sequences, avoiding silent mutations caused by deletion of non-coding sequences. (FIG. 6 shows the intron/exon structure for the human AIB1 gene). These cells were microinjected into mouse embryos leading to the deletion of the mouse AIB1 gene in the germ line of a transgenic mouse. The mouse AIB1 gene was mapped and isolated by the following method: The primers AIB/mEST F1 (5'-TCCTTTTCCCAGCAGCAGTTTG-3'; SEQ. I.D. 10) and AIB1/mEST R1 (5'ATGCCAGACATGGGCATGGG-3' SEQ. I.D. 11) were used to screen a mouse Bacterial Artificial Chromosome (BAC) library and to isolate a mouse BAC (designated 195H10). This BAC was assigned to mouse chromosome 2 by fluorescence in situ hybridization (FISH). This region is the mouse equivalent of the portion of human chromosome 20 which carries AIB1.
[0119]To map the structure of the gene, first the structure of the human AIB1 gene was determined by polymerase chain reaction of a human genomic DNA clone containing AIB1 using standard methods (Genomics 1995 Jan. 20; 25(2):501-506) and then the sequences of the intron exon boundaries were determined (FIG. 4). Based on this information, the corresponding regions of the mouse BAC were sequenced. The structure of the mouse gene corresponds closely to that of the human gene (FIG. 4). This information localizes the coding regions of the mouse AIB1 gene so that a targeting vector can be constructed to remove these regions from mouse embryonic stem cells. These cells can be then injected into mouse embryos leading to deletion of the mouse AIB1 gene in the germ line of a transgenic mouse. The methods of creating deletion mutations by using a targeting vector have been described in Cell (Thomas and Capecch, Cell 51(3):503-512, 1987).
[0120]References and patents referred to herein are incorporated by reference.
[0121]The above examples are provided by way of illustration only and are in no way intended to limit the scope of the invention. One of skill in the art will see that the invention may be modified in various ways without departing from the spirit or principle of the invention. We claim all such modifications.
Sequence CWU
1
1216835DNAHomo sapiensCDS(201)..(4463) 1cggcggcggc tgcggcttag tcggtggcgg
ccggcggcgg ctgcgggctg agcggcgagt 60ttccgattta aagctgagct gcgaggaaaa
tggcggcggg aggatcaaaa tacttgctgg 120atggtggact cagagaccaa taaaaataaa
ctgcttgaac atcctttgac tggttagcca 180gttgctgatg tatattcaag atg agt gga
tta gga gaa aac ttg gat cca ctg 233Met Ser Gly Leu Gly Glu Asn Leu Asp
Pro Leu1 5 10gcc agt gat tca cga aaa cgc
aaa ttg cca tgt gat act cca gga caa 281Ala Ser Asp Ser Arg Lys Arg
Lys Leu Pro Cys Asp Thr Pro Gly Gln15 20
25ggt ctt acc tgc agt ggt gaa aaa cgg aga cgg gag cag gaa agt aaa
329Gly Leu Thr Cys Ser Gly Glu Lys Arg Arg Arg Glu Gln Glu Ser Lys30
35 40tat att gaa gaa ttg gct gag ctg ata tct
gcc aat ctt agt gat att 377Tyr Ile Glu Glu Leu Ala Glu Leu Ile Ser
Ala Asn Leu Ser Asp Ile45 50 55gac aat
ttc aat gtc aaa cca gat aaa tgt gcg att tta aag gaa aca 425Asp Asn
Phe Asn Val Lys Pro Asp Lys Cys Ala Ile Leu Lys Glu Thr60
65 70 75gta aga cag ata cgt caa ata
aaa gag caa gga aaa act att tcc aat 473Val Arg Gln Ile Arg Gln Ile
Lys Glu Gln Gly Lys Thr Ile Ser Asn80 85
90gat gat gat gtt caa aaa gcc gat gta tct tct aca ggg cag gga gtt
521Asp Asp Asp Val Gln Lys Ala Asp Val Ser Ser Thr Gly Gln Gly Val95
100 105att gat aaa gac tcc tta gga ccg ctt tta
ctt cag gca ttg gat ggt 569Ile Asp Lys Asp Ser Leu Gly Pro Leu Leu
Leu Gln Ala Leu Asp Gly110 115 120ttc cta
ttt gtg gtg aat cga gac gga aac att gta ttt gta tca gaa 617Phe Leu
Phe Val Val Asn Arg Asp Gly Asn Ile Val Phe Val Ser Glu125
130 135aat gtc aca caa tac ctg caa tat aag caa gag gac
ctg gtt aac aca 665Asn Val Thr Gln Tyr Leu Gln Tyr Lys Gln Glu Asp
Leu Val Asn Thr140 145 150
155agt gtt tac aat atc tta cat gaa gaa gac aga aag gat ttt ctt aag
713Ser Val Tyr Asn Ile Leu His Glu Glu Asp Arg Lys Asp Phe Leu Lys160
165 170aat tta cca aaa tct aca gtt aat gga
gtt tcc tgg aca aat gag acc 761Asn Leu Pro Lys Ser Thr Val Asn Gly
Val Ser Trp Thr Asn Glu Thr175 180 185caa
aga caa aaa agc cat aca ttt aat tgc cgt atg ttg atg aaa aca 809Gln
Arg Gln Lys Ser His Thr Phe Asn Cys Arg Met Leu Met Lys Thr190
195 200cca cat gat att ctg gaa gac ata aac gcc agt
cct gaa atg cgc cag 857Pro His Asp Ile Leu Glu Asp Ile Asn Ala Ser
Pro Glu Met Arg Gln205 210 215aga tat gaa
aca atg cag tgc ttt gcc ctg tct cag cca cga gct atg 905Arg Tyr Glu
Thr Met Gln Cys Phe Ala Leu Ser Gln Pro Arg Ala Met220
225 230 235atg gag gaa ggg gaa gat ttg
caa tct tgt atg atc tgt gtg gca cgc 953Met Glu Glu Gly Glu Asp Leu
Gln Ser Cys Met Ile Cys Val Ala Arg240 245
250cgc att act aca gga gaa aga aca ttt cca tca aac cct gag agc ttt
1001Arg Ile Thr Thr Gly Glu Arg Thr Phe Pro Ser Asn Pro Glu Ser Phe255
260 265att acc aga cat gat ctt tca gga aag
gtt gtc aat ata gat aca aat 1049Ile Thr Arg His Asp Leu Ser Gly Lys
Val Val Asn Ile Asp Thr Asn270 275 280tca
ctg aga tcc tcc atg agg cct ggc ttt gaa gat ata atc cga agg 1097Ser
Leu Arg Ser Ser Met Arg Pro Gly Phe Glu Asp Ile Ile Arg Arg285
290 295tgt att cag aga ttt ttt agt cta aat gat ggg
cag tca tgg tcc cag 1145Cys Ile Gln Arg Phe Phe Ser Leu Asn Asp Gly
Gln Ser Trp Ser Gln300 305 310
315aaa cgt cac tat caa gaa gct tat ctt aat ggc cat gca gaa acc cca
1193Lys Arg His Tyr Gln Glu Ala Tyr Leu Asn Gly His Ala Glu Thr Pro320
325 330gta tat cga ttc tcg ttg gct gat gga
act ata gtg act gca cag aca 1241Val Tyr Arg Phe Ser Leu Ala Asp Gly
Thr Ile Val Thr Ala Gln Thr335 340 345aaa
agc aaa ctc ttc cga aat cct gta aca aat gat cga cat ggc ttt 1289Lys
Ser Lys Leu Phe Arg Asn Pro Val Thr Asn Asp Arg His Gly Phe350
355 360gtc tca acc cac ttc ctt cag aga gaa cag aat
gga tat aga cca aac 1337Val Ser Thr His Phe Leu Gln Arg Glu Gln Asn
Gly Tyr Arg Pro Asn365 370 375cca aat cct
gtt gga caa ggg att aga cca cct atg gct gga tgc aac 1385Pro Asn Pro
Val Gly Gln Gly Ile Arg Pro Pro Met Ala Gly Cys Asn380
385 390 395agt tcg gta ggc ggc atg agt
atg tcg cca aac caa ggc tta cag atg 1433Ser Ser Val Gly Gly Met Ser
Met Ser Pro Asn Gln Gly Leu Gln Met400 405
410ccg agc agc agg gcc tat ggc ttg gca gac cct agc acc aca ggg cag
1481Pro Ser Ser Arg Ala Tyr Gly Leu Ala Asp Pro Ser Thr Thr Gly Gln415
420 425atg agt gga gct agg tat ggg ggt tcc
agt aac ata gct tca ttg acc 1529Met Ser Gly Ala Arg Tyr Gly Gly Ser
Ser Asn Ile Ala Ser Leu Thr430 435 440cct
ggg cca ggc atg caa tca cca tct tcc tac cag aac aac aac tat 1577Pro
Gly Pro Gly Met Gln Ser Pro Ser Ser Tyr Gln Asn Asn Asn Tyr445
450 455ggg ctc aac atg agt agc ccc cca cat ggg agt
cct ggt ctt gcc cca 1625Gly Leu Asn Met Ser Ser Pro Pro His Gly Ser
Pro Gly Leu Ala Pro460 465 470
475aac cag cag aat atc atg att tct cct cgt aat cgt ggg agt cca aag
1673Asn Gln Gln Asn Ile Met Ile Ser Pro Arg Asn Arg Gly Ser Pro Lys480
485 490ata gcc tca cat cag ttt tct cct gtt
gca ggt gtg cac tct ccc atg 1721Ile Ala Ser His Gln Phe Ser Pro Val
Ala Gly Val His Ser Pro Met495 500 505gca
tct tct ggc aat act ggg aac cac agc ttt tcc agc agc tct ctc 1769Ala
Ser Ser Gly Asn Thr Gly Asn His Ser Phe Ser Ser Ser Ser Leu510
515 520agt gcc ctg caa gcc atc agt gaa ggt gtg ggg
act tcc ctt tta tct 1817Ser Ala Leu Gln Ala Ile Ser Glu Gly Val Gly
Thr Ser Leu Leu Ser525 530 535act ctg tca
tca cca ggc ccc aaa ttg gat aac tct ccc aat atg aat 1865Thr Leu Ser
Ser Pro Gly Pro Lys Leu Asp Asn Ser Pro Asn Met Asn540
545 550 555att acc caa cca agt aaa gta
agc aat cag gat tcc aag agt cct ctg 1913Ile Thr Gln Pro Ser Lys Val
Ser Asn Gln Asp Ser Lys Ser Pro Leu560 565
570ggc ttt tat tgc gac caa aat cca gtg gag agt tca atg tgt cag tca
1961Gly Phe Tyr Cys Asp Gln Asn Pro Val Glu Ser Ser Met Cys Gln Ser575
580 585aat agc aga gat cac ctc agt gac aaa
gaa agt aag gag agc agt gtt 2009Asn Ser Arg Asp His Leu Ser Asp Lys
Glu Ser Lys Glu Ser Ser Val590 595 600gag
ggg gca gag aat caa agg ggt cct ttg gaa agc aaa ggt cat aaa 2057Glu
Gly Ala Glu Asn Gln Arg Gly Pro Leu Glu Ser Lys Gly His Lys605
610 615aaa tta ctg cag tta ctt acc tgt tct tct gat
gac cgg ggt cat tcc 2105Lys Leu Leu Gln Leu Leu Thr Cys Ser Ser Asp
Asp Arg Gly His Ser620 625 630
635tcc ttg acc aac tcc ccc cta gat tca agt tgt aaa gaa tct tct gtt
2153Ser Leu Thr Asn Ser Pro Leu Asp Ser Ser Cys Lys Glu Ser Ser Val640
645 650agt gtc acc agc ccc tct gga gtc tcc
tcc tct aca tct gga gga gta 2201Ser Val Thr Ser Pro Ser Gly Val Ser
Ser Ser Thr Ser Gly Gly Val655 660 665tcc
tct aca tcc aat atg cat ggg tca ctg tta caa gag aag cac cgg 2249Ser
Ser Thr Ser Asn Met His Gly Ser Leu Leu Gln Glu Lys His Arg670
675 680att ttg cac aag ttg ctg cag aat ggg aat tca
cca gct gag gta gcc 2297Ile Leu His Lys Leu Leu Gln Asn Gly Asn Ser
Pro Ala Glu Val Ala685 690 695aag att act
gca gaa gcc act ggg aaa gac acc agc agt ata act tct 2345Lys Ile Thr
Ala Glu Ala Thr Gly Lys Asp Thr Ser Ser Ile Thr Ser700
705 710 715tgt ggg gac gga aat gtt gtc
aag cag gag cag cta agt cct aag aag 2393Cys Gly Asp Gly Asn Val Val
Lys Gln Glu Gln Leu Ser Pro Lys Lys720 725
730aag gag aat aat gca ctt ctt aga tac ctg ctg gac agg gat gat cct
2441Lys Glu Asn Asn Ala Leu Leu Arg Tyr Leu Leu Asp Arg Asp Asp Pro735
740 745agt gat gca ctc tct aaa gaa cta cag
ccc caa gtg gaa gga gtg gat 2489Ser Asp Ala Leu Ser Lys Glu Leu Gln
Pro Gln Val Glu Gly Val Asp750 755 760aat
aaa atg agt cag tgc acc agc tcc acc att cct agc tca agt caa 2537Asn
Lys Met Ser Gln Cys Thr Ser Ser Thr Ile Pro Ser Ser Ser Gln765
770 775gag aaa gac cct aaa att aag aca gag aca agt
gaa gag gga tct gga 2585Glu Lys Asp Pro Lys Ile Lys Thr Glu Thr Ser
Glu Glu Gly Ser Gly780 785 790
795gac ttg gat aat cta gat gct att ctt ggt gat ctg act agt tct gac
2633Asp Leu Asp Asn Leu Asp Ala Ile Leu Gly Asp Leu Thr Ser Ser Asp800
805 810ttt tac aat aat tcc ata tcc tca aat
ggt agt cat ctg ggg act aag 2681Phe Tyr Asn Asn Ser Ile Ser Ser Asn
Gly Ser His Leu Gly Thr Lys815 820 825caa
cag gtg ttt caa gga act aat tct ctg ggt ttg aaa agt tca cag 2729Gln
Gln Val Phe Gln Gly Thr Asn Ser Leu Gly Leu Lys Ser Ser Gln830
835 840tct gtg cag tct att cgt cct cca tat aac cga
gca gtg tct ctg gat 2777Ser Val Gln Ser Ile Arg Pro Pro Tyr Asn Arg
Ala Val Ser Leu Asp845 850 855agc cct gtt
tct gtt ggc tca agt cct cca gta aaa aat atc agt gct 2825Ser Pro Val
Ser Val Gly Ser Ser Pro Pro Val Lys Asn Ile Ser Ala860
865 870 875ttc ccc atg tta cca aag caa
ccc atg ttg ggt ggg aat cca aga atg 2873Phe Pro Met Leu Pro Lys Gln
Pro Met Leu Gly Gly Asn Pro Arg Met880 885
890atg gat agt cag gaa aat tat ggc tca agt atg ggt ggg cca aac cga
2921Met Asp Ser Gln Glu Asn Tyr Gly Ser Ser Met Gly Gly Pro Asn Arg895
900 905aat gtg act gtg act cag act cct tcc
tca gga gac tgg ggc tta cca 2969Asn Val Thr Val Thr Gln Thr Pro Ser
Ser Gly Asp Trp Gly Leu Pro910 915 920aac
tca aag gcc ggc aga atg gaa cct atg aat tca aac tcc atg gga 3017Asn
Ser Lys Ala Gly Arg Met Glu Pro Met Asn Ser Asn Ser Met Gly925
930 935aga cca gga gga gat tat aat act tct tta ccc
aga cct gca ctg ggt 3065Arg Pro Gly Gly Asp Tyr Asn Thr Ser Leu Pro
Arg Pro Ala Leu Gly940 945 950
955ggc tct att ccc aca ttg cct ctt cgg tct aat agc ata cca ggt gcg
3113Gly Ser Ile Pro Thr Leu Pro Leu Arg Ser Asn Ser Ile Pro Gly Ala960
965 970aga cca gta ttg caa cag cag cag cag
atg ctt caa atg agg cct ggt 3161Arg Pro Val Leu Gln Gln Gln Gln Gln
Met Leu Gln Met Arg Pro Gly975 980 985gaa
atc ccc atg gga atg ggg gct aat ccc tat ggc caa gca gca gca 3209Glu
Ile Pro Met Gly Met Gly Ala Asn Pro Tyr Gly Gln Ala Ala Ala990
995 1000tct aac caa ctg ggt tcc tgg ccc gat ggc atg
ttg tcc atg gaa caa 3257Ser Asn Gln Leu Gly Ser Trp Pro Asp Gly Met
Leu Ser Met Glu Gln1005 1010 1015gtt tct
cat ggc act caa aat agg cct ctt ctt agg aat tcc ctg gat 3305Val Ser
His Gly Thr Gln Asn Arg Pro Leu Leu Arg Asn Ser Leu Asp1020
1025 1030 1035gat ctt gtt ggg cca cct tcc
aac ctg gaa ggc cag agt gac gaa aga 3353Asp Leu Val Gly Pro Pro Ser
Asn Leu Glu Gly Gln Ser Asp Glu Arg1040 1045
1050gca tta ttg gac cag ctg cac act ctt ctc agc aac aca gat gcc aca
3401Ala Leu Leu Asp Gln Leu His Thr Leu Leu Ser Asn Thr Asp Ala Thr1055
1060 1065ggc ctg gaa gaa att gac aga gct ttg
ggc att cct gaa ctt gtc aat 3449Gly Leu Glu Glu Ile Asp Arg Ala Leu
Gly Ile Pro Glu Leu Val Asn1070 1075
1080cag gga cag gca tta gag ccc aaa cag gat gct ttc caa ggc caa gaa
3497Gln Gly Gln Ala Leu Glu Pro Lys Gln Asp Ala Phe Gln Gly Gln Glu1085
1090 1095gca gca gta atg atg gat cag aag gca
gga tta tat gga cag aca tac 3545Ala Ala Val Met Met Asp Gln Lys Ala
Gly Leu Tyr Gly Gln Thr Tyr1100 1105 1110
1115cca gca cag ggg cct cca atg caa gga ggc ttt cat ctt cag
gga caa 3593Pro Ala Gln Gly Pro Pro Met Gln Gly Gly Phe His Leu Gln
Gly Gln1120 1125 1130tca cca tct ttt aac
tct atg atg aat cag atg aac cag caa ggc aat 3641Ser Pro Ser Phe Asn
Ser Met Met Asn Gln Met Asn Gln Gln Gly Asn1135 1140
1145ttt cct ctc caa gga atg cac cca cga gcc aac atc atg aga ccc
cgg 3689Phe Pro Leu Gln Gly Met His Pro Arg Ala Asn Ile Met Arg Pro
Arg1150 1155 1160aca aac acc ccc aag caa
ctt aga atg cag ctt cag cag agg ctg cag 3737Thr Asn Thr Pro Lys Gln
Leu Arg Met Gln Leu Gln Gln Arg Leu Gln1165 1170
1175ggc cag cag ttt ttg aat cag agc cga cag gca ctt gaa ttg aaa atg
3785Gly Gln Gln Phe Leu Asn Gln Ser Arg Gln Ala Leu Glu Leu Lys
Met1180 1185 1190 1195gaa aac
cct act gct ggt ggt gct gcg gtg atg agg cct atg atg cag 3833Glu Asn
Pro Thr Ala Gly Gly Ala Ala Val Met Arg Pro Met Met Gln1200
1205 1210ccc cag cag ggt ttt ctt aat gct caa atg gtc gcc
caa cgc agc aga 3881Pro Gln Gln Gly Phe Leu Asn Ala Gln Met Val Ala
Gln Arg Ser Arg1215 1220 1225gag ctg cta
agt cat cac ttc cga caa cag agg gtg gct atg atg atg 3929Glu Leu Leu
Ser His His Phe Arg Gln Gln Arg Val Ala Met Met Met1230
1235 1240cag cag cag cag cag cag caa cag cag cag cag cag
cag cag cag cag 3977Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln
Gln Gln Gln Gln1245 1250 1255caa cag caa
cag caa cag caa cag cag caa cag cag caa acc cag gcc 4025Gln Gln Gln
Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Thr Gln Ala1260
1265 1270 1275ttc agc cca cct cct aat gtg
act gct tcc ccc agc atg gat ggg ctt 4073Phe Ser Pro Pro Pro Asn Val
Thr Ala Ser Pro Ser Met Asp Gly Leu1280 1285
1290ttg gca gga ccc aca atg cca caa gct cct ccg caa cag ttt cca tat
4121Leu Ala Gly Pro Thr Met Pro Gln Ala Pro Pro Gln Gln Phe Pro Tyr1295
1300 1305caa cca aat tat gga atg gga caa caa
cca gat cca gcc ttt ggt cga 4169Gln Pro Asn Tyr Gly Met Gly Gln Gln
Pro Asp Pro Ala Phe Gly Arg1310 1315
1320gtg tct agt cct ccc aat gca atg atg tcg tca aga atg ggt ccc tcc
4217Val Ser Ser Pro Pro Asn Ala Met Met Ser Ser Arg Met Gly Pro Ser1325
1330 1335cag aat ccc atg atg caa cac ccg cag
gct gca tcc atc tat cag tcc 4265Gln Asn Pro Met Met Gln His Pro Gln
Ala Ala Ser Ile Tyr Gln Ser1340 1345 1350
1355tca gaa atg aag ggc tgg cca tca gga aat ttg gcc agg aac
agc tcc 4313Ser Glu Met Lys Gly Trp Pro Ser Gly Asn Leu Ala Arg Asn
Ser Ser1360 1365 1370ttt tcc cag cag cag
ttt gcc cac cag ggg aat cct gca gtg tat agt 4361Phe Ser Gln Gln Gln
Phe Ala His Gln Gly Asn Pro Ala Val Tyr Ser1375 1380
1385atg gtg cac atg aat ggc agc agt ggt cac atg gga cag atg aac
atg 4409Met Val His Met Asn Gly Ser Ser Gly His Met Gly Gln Met Asn
Met1390 1395 1400aac ccc atg ccc atg tct
ggc atg cct atg ggt cct gat cag aaa tac 4457Asn Pro Met Pro Met Ser
Gly Met Pro Met Gly Pro Asp Gln Lys Tyr1405 1410
1415tgc tga catctctgca ccaggacctc ttaaggaaac cactgtacaa atgacactgc
4513Cys1420actaggatta ttgggaagga atcattgttc caggcatcca tcttggaaga
aaggaccagc 4573tttgagctcc atcaagggta ttttaagtga tgtcatttga gcaggactgg
attttaagcc 4633gaagggcaat atctacgtgt ttttcccccc tccttctgct gtgtatcatg
gtgttcaaaa 4693cagaaatgtt ttttggcatt ccacctccta gggatataat tctggagaca
tggagtgtta 4753ctgatcataa aacttttgtg tcactttttt ctgccttgct agccaaaatc
tcttaaatac 4813acgtaggtgg gccagagaac attggaagaa tcaagagaga ttagaatatc
tggtttctct 4873agttgcagta ttggacaaag agcatagtcc cagccttcag gtgtagtagt
tctgtgttga 4933ccctttgtcc agtggaattg gtgattctga attgtccttt actaatggtg
ttgagttgct 4993ctgtccctat tatttgccct aggctttctc ctaatgaagg ttttcatttg
ccattcatgt 5053cctgtaatac ttcacctcca ggaactgtca tggatgtcca aatggctttg
cagaaaggaa 5113atgagatgac agtatttaat cgcagcagta gcaaactttt cacatgctaa
tgtgcagctg 5173agtgcacttt atttaaaaag aatggataaa tgcaatattc ttgaggtctt
gagggaatag 5233tgaaacacat tcctggtttt tgcctacact tacgtgttag acaagaacta
tgattttttt 5293tttaaagtac tggtgtcacc ctttgcctat atggtagagc aataatgctt
tttaaaaata 5353aacttctgaa aacccaaggc caggtactgc attctgaatc agaatctcgc
agtgtttctg 5413tgaatagatt tttttgtaaa tatgaccttt aagatattgt attatgtaaa
atatgtatat 5473accttttttt gtaggtcaca acaactcatt tttacagagt ttgtgaagct
aaatatttaa 5533cattgttgat ttcagtaagc tgtgtggtga ggctaccagt ggaagagaca
tcccttgact 5593tttgtggcct gggggagggg tagtgctcca cagcttttcc ttccccaccc
cccagcctta 5653gatgcctcgc tcttttcaat ctcttaatct aaatgctttt taaagagatt
atttgtttag 5713atgtaggcat tttaattttt taaaaattcc tctaccagaa ctaagcactt
tgttaatttg 5773gggggaaaga atagatatgg ggaaataaac ttaaaaaaaa atcaggaatt
taaaaaaacg 5833agcaatttga agagaatctt ttggatttta agcagtccga aataatagca
attcatgggc 5893tgtgtgtgtg tgtgtatgtg tgtgtgtgtg tgtgtatgtt taattatgtt
accttttcat 5953cccctttagg agcgttttca gattttggtt gctaagacct gaatcccata
ttgagatctc 6013gagtagaatc cttggtgtgg tttctggtgt ctgctcagct gtcccctcat
tctactaatg 6073tgatgctttc attatgtccc tgtggattag aatagtgtca gttatttctt
aagtaactca 6133gtacccagaa cagccagttt tactgtgatt cagagccaca gtctaactga
gcacctttta 6193aacccctccc tcttctgccc cctaccactt ttctgctgtt gcctctcttt
gacacctgtt 6253ttagtcagtt gggaggaagg gaaaaatcaa gtttaattcc ctttatctgg
gttaattcat 6313ttggttcaaa tagttgacgg aattgggttt ctgaatgtct gtgaatttca
gaggtctctg 6373ctagccttgg tatcattttc tagcaataac tgagagccag ttaattttaa
gaatttcaca 6433catttagcca atctttctag atgtctctga aggtaagatc atttaatatc
tttgatatgc 6493ttacgagtaa gtgaatcctg attatttcca gacccaccac cagagtggat
cttattttca 6553aagcagtata gacaattatg agtttgccct ctttccccta ccaagttcaa
aatatatcta 6613agaaagattg taaatccgaa aacttccatt gtagtggcct gtgcttttca
gatagtatac 6673tctcctgttt ggagacagag gaagaaccag gtcagtctgt ctctttttca
gctcaattgt 6733atctgaccct tctttaagtt atgtgtgtgg ggagaaatag aatggtgctc
ttatctttct 6793tgactttaaa aaaattatta aaaacaaaaa aaaaaaaaaa aa
68352186PRTHomo sapiens 2Leu Leu Gln Ala Leu Asp Gly Phe Leu
Phe Val Val Asn Arg Asp Gly1 5 10
15Asn Ile Val Phe Val Ser Glu Asn Val Thr Gln Tyr Leu Gln Tyr
Lys20 25 30Gln Glu Asp Leu Val Asn Thr
Ser Val Tyr Asn Ile Leu His Glu Glu35 40
45Asp Arg Lys Asp Phe Leu Lys Asn Leu Pro Lys Ser Thr Val Asn Gly50
55 60Val Ser Trp Thr Asn Glu Thr Gln Arg Gln
Lys Ser His Thr Phe Asn65 70 75
80Cys Arg Met Leu Met Lys Thr Pro His Asp Ile Leu Glu Asp Ile
Asn85 90 95Ala Ser Pro Glu Met Arg Gln
Arg Tyr Glu Thr Met Gln Cys Phe Ala100 105
110Leu Ser Gln Pro Arg Ala Met Met Glu Glu Gly Glu Asp Leu Gln Ser115
120 125Cys Met Ile Cys Val Ala Arg Arg Ile
Thr Thr Gly Glu Arg Thr Phe130 135 140Pro
Ser Asn Pro Glu Ser Phe Ile Thr Arg His Asp Leu Ser Gly Lys145
150 155 160Val Val Asn Ile Asp Thr
Asn Ser Leu Arg Ser Ser Met Arg Pro Gly165 170
175Phe Glu Asp Ile Ile Arg Arg Cys Ile Gln180
185373PRTHomo sapiens 3Arg Lys Arg Lys Leu Pro Cys Asp Thr Pro Gly Gln
Gly Leu Thr Cys1 5 10
15Ser Gly Glu Lys Arg Arg Arg Glu Gln Glu Ser Lys Tyr Ile Glu Glu20
25 30Leu Ala Glu Leu Ile Ser Ala Asn Leu Ser
Asp Ile Asp Asn Phe Asn35 40 45Val Lys
Pro Asp Lys Cys Ala Ile Leu Lys Glu Thr Val Arg Gln Ile50
55 60Arg Gln Ile Lys Glu Gln Gly Lys Thr65
7041420PRTHomo sapiens 4Met Ser Gly Leu Gly Glu Asn Leu Asp Pro Leu
Ala Ser Asp Ser Arg1 5 10
15Lys Arg Lys Leu Pro Cys Asp Thr Pro Gly Gln Gly Leu Thr Cys Ser20
25 30Gly Glu Lys Arg Arg Arg Glu Gln Glu Ser
Lys Tyr Ile Glu Glu Leu35 40 45Ala Glu
Leu Ile Ser Ala Asn Leu Ser Asp Ile Asp Asn Phe Asn Val50
55 60Lys Pro Asp Lys Cys Ala Ile Leu Lys Glu Thr Val
Arg Gln Ile Arg65 70 75
80Gln Ile Lys Glu Gln Gly Lys Thr Ile Ser Asn Asp Asp Asp Val Gln85
90 95Lys Ala Asp Val Ser Ser Thr Gly Gln Gly
Val Ile Asp Lys Asp Ser100 105 110Leu Gly
Pro Leu Leu Leu Gln Ala Leu Asp Gly Phe Leu Phe Val Val115
120 125Asn Arg Asp Gly Asn Ile Val Phe Val Ser Glu Asn
Val Thr Gln Tyr130 135 140Leu Gln Tyr Lys
Gln Glu Asp Leu Val Asn Thr Ser Val Tyr Asn Ile145 150
155 160Leu His Glu Glu Asp Arg Lys Asp Phe
Leu Lys Asn Leu Pro Lys Ser165 170 175Thr
Val Asn Gly Val Ser Trp Thr Asn Glu Thr Gln Arg Gln Lys Ser180
185 190His Thr Phe Asn Cys Arg Met Leu Met Lys Thr
Pro His Asp Ile Leu195 200 205Glu Asp Ile
Asn Ala Ser Pro Glu Met Arg Gln Arg Tyr Glu Thr Met210
215 220Gln Cys Phe Ala Leu Ser Gln Pro Arg Ala Met Met
Glu Glu Gly Glu225 230 235
240Asp Leu Gln Ser Cys Met Ile Cys Val Ala Arg Arg Ile Thr Thr Gly245
250 255Glu Arg Thr Phe Pro Ser Asn Pro Glu
Ser Phe Ile Thr Arg His Asp260 265 270Leu
Ser Gly Lys Val Val Asn Ile Asp Thr Asn Ser Leu Arg Ser Ser275
280 285Met Arg Pro Gly Phe Glu Asp Ile Ile Arg Arg
Cys Ile Gln Arg Phe290 295 300Phe Ser Leu
Asn Asp Gly Gln Ser Trp Ser Gln Lys Arg His Tyr Gln305
310 315 320Glu Ala Tyr Leu Asn Gly His
Ala Glu Thr Pro Val Tyr Arg Phe Ser325 330
335Leu Ala Asp Gly Thr Ile Val Thr Ala Gln Thr Lys Ser Lys Leu Phe340
345 350Arg Asn Pro Val Thr Asn Asp Arg His
Gly Phe Val Ser Thr His Phe355 360 365Leu
Gln Arg Glu Gln Asn Gly Tyr Arg Pro Asn Pro Asn Pro Val Gly370
375 380Gln Gly Ile Arg Pro Pro Met Ala Gly Cys Asn
Ser Ser Val Gly Gly385 390 395
400Met Ser Met Ser Pro Asn Gln Gly Leu Gln Met Pro Ser Ser Arg
Ala405 410 415Tyr Gly Leu Ala Asp Pro Ser
Thr Thr Gly Gln Met Ser Gly Ala Arg420 425
430Tyr Gly Gly Ser Ser Asn Ile Ala Ser Leu Thr Pro Gly Pro Gly Met435
440 445Gln Ser Pro Ser Ser Tyr Gln Asn Asn
Asn Tyr Gly Leu Asn Met Ser450 455 460Ser
Pro Pro His Gly Ser Pro Gly Leu Ala Pro Asn Gln Gln Asn Ile465
470 475 480Met Ile Ser Pro Arg Asn
Arg Gly Ser Pro Lys Ile Ala Ser His Gln485 490
495Phe Ser Pro Val Ala Gly Val His Ser Pro Met Ala Ser Ser Gly
Asn500 505 510Thr Gly Asn His Ser Phe Ser
Ser Ser Ser Leu Ser Ala Leu Gln Ala515 520
525Ile Ser Glu Gly Val Gly Thr Ser Leu Leu Ser Thr Leu Ser Ser Pro530
535 540Gly Pro Lys Leu Asp Asn Ser Pro Asn
Met Asn Ile Thr Gln Pro Ser545 550 555
560Lys Val Ser Asn Gln Asp Ser Lys Ser Pro Leu Gly Phe Tyr
Cys Asp565 570 575Gln Asn Pro Val Glu Ser
Ser Met Cys Gln Ser Asn Ser Arg Asp His580 585
590Leu Ser Asp Lys Glu Ser Lys Glu Ser Ser Val Glu Gly Ala Glu
Asn595 600 605Gln Arg Gly Pro Leu Glu Ser
Lys Gly His Lys Lys Leu Leu Gln Leu610 615
620Leu Thr Cys Ser Ser Asp Asp Arg Gly His Ser Ser Leu Thr Asn Ser625
630 635 640Pro Leu Asp Ser
Ser Cys Lys Glu Ser Ser Val Ser Val Thr Ser Pro645 650
655Ser Gly Val Ser Ser Ser Thr Ser Gly Gly Val Ser Ser Thr
Ser Asn660 665 670Met His Gly Ser Leu Leu
Gln Glu Lys His Arg Ile Leu His Lys Leu675 680
685Leu Gln Asn Gly Asn Ser Pro Ala Glu Val Ala Lys Ile Thr Ala
Glu690 695 700Ala Thr Gly Lys Asp Thr Ser
Ser Ile Thr Ser Cys Gly Asp Gly Asn705 710
715 720Val Val Lys Gln Glu Gln Leu Ser Pro Lys Lys Lys
Glu Asn Asn Ala725 730 735Leu Leu Arg Tyr
Leu Leu Asp Arg Asp Asp Pro Ser Asp Ala Leu Ser740 745
750Lys Glu Leu Gln Pro Gln Val Glu Gly Val Asp Asn Lys Met
Ser Gln755 760 765Cys Thr Ser Ser Thr Ile
Pro Ser Ser Ser Gln Glu Lys Asp Pro Lys770 775
780Ile Lys Thr Glu Thr Ser Glu Glu Gly Ser Gly Asp Leu Asp Asn
Leu785 790 795 800Asp Ala
Ile Leu Gly Asp Leu Thr Ser Ser Asp Phe Tyr Asn Asn Ser805
810 815Ile Ser Ser Asn Gly Ser His Leu Gly Thr Lys Gln
Gln Val Phe Gln820 825 830Gly Thr Asn Ser
Leu Gly Leu Lys Ser Ser Gln Ser Val Gln Ser Ile835 840
845Arg Pro Pro Tyr Asn Arg Ala Val Ser Leu Asp Ser Pro Val
Ser Val850 855 860Gly Ser Ser Pro Pro Val
Lys Asn Ile Ser Ala Phe Pro Met Leu Pro865 870
875 880Lys Gln Pro Met Leu Gly Gly Asn Pro Arg Met
Met Asp Ser Gln Glu885 890 895Asn Tyr Gly
Ser Ser Met Gly Gly Pro Asn Arg Asn Val Thr Val Thr900
905 910Gln Thr Pro Ser Ser Gly Asp Trp Gly Leu Pro Asn
Ser Lys Ala Gly915 920 925Arg Met Glu Pro
Met Asn Ser Asn Ser Met Gly Arg Pro Gly Gly Asp930 935
940Tyr Asn Thr Ser Leu Pro Arg Pro Ala Leu Gly Gly Ser Ile
Pro Thr945 950 955 960Leu
Pro Leu Arg Ser Asn Ser Ile Pro Gly Ala Arg Pro Val Leu Gln965
970 975Gln Gln Gln Gln Met Leu Gln Met Arg Pro Gly
Glu Ile Pro Met Gly980 985 990Met Gly Ala
Asn Pro Tyr Gly Gln Ala Ala Ala Ser Asn Gln Leu Gly995
1000 1005Ser Trp Pro Asp Gly Met Leu Ser Met Glu Gln Val
Ser His Gly Thr1010 1015 1020Gln Asn Arg
Pro Leu Leu Arg Asn Ser Leu Asp Asp Leu Val Gly Pro1025
1030 1035 1040Pro Ser Asn Leu Glu Gly Gln
Ser Asp Glu Arg Ala Leu Leu Asp Gln1045 1050
1055Leu His Thr Leu Leu Ser Asn Thr Asp Ala Thr Gly Leu Glu Glu Ile1060
1065 1070Asp Arg Ala Leu Gly Ile Pro Glu Leu
Val Asn Gln Gly Gln Ala Leu1075 1080
1085Glu Pro Lys Gln Asp Ala Phe Gln Gly Gln Glu Ala Ala Val Met Met1090
1095 1100Asp Gln Lys Ala Gly Leu Tyr Gly Gln
Thr Tyr Pro Ala Gln Gly Pro1105 1110 1115
1120Pro Met Gln Gly Gly Phe His Leu Gln Gly Gln Ser Pro Ser
Phe Asn1125 1130 1135Ser Met Met Asn Gln
Met Asn Gln Gln Gly Asn Phe Pro Leu Gln Gly1140 1145
1150Met His Pro Arg Ala Asn Ile Met Arg Pro Arg Thr Asn Thr Pro
Lys1155 1160 1165Gln Leu Arg Met Gln Leu
Gln Gln Arg Leu Gln Gly Gln Gln Phe Leu1170 1175
1180Asn Gln Ser Arg Gln Ala Leu Glu Leu Lys Met Glu Asn Pro Thr
Ala1185 1190 1195 1200Gly Gly
Ala Ala Val Met Arg Pro Met Met Gln Pro Gln Gln Gly Phe1205
1210 1215Leu Asn Ala Gln Met Val Ala Gln Arg Ser Arg Glu
Leu Leu Ser His1220 1225 1230His Phe Arg
Gln Gln Arg Val Ala Met Met Met Gln Gln Gln Gln Gln1235
1240 1245Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln
Gln Gln Gln Gln1250 1255 1260Gln Gln Gln
Gln Gln Gln Gln Gln Thr Gln Ala Phe Ser Pro Pro Pro1265
1270 1275 1280Asn Val Thr Ala Ser Pro Ser
Met Asp Gly Leu Leu Ala Gly Pro Thr1285 1290
1295Met Pro Gln Ala Pro Pro Gln Gln Phe Pro Tyr Gln Pro Asn Tyr Gly1300
1305 1310Met Gly Gln Gln Pro Asp Pro Ala Phe
Gly Arg Val Ser Ser Pro Pro1315 1320
1325Asn Ala Met Met Ser Ser Arg Met Gly Pro Ser Gln Asn Pro Met Met1330
1335 1340Gln His Pro Gln Ala Ala Ser Ile Tyr
Gln Ser Ser Glu Met Lys Gly1345 1350 1355
1360Trp Pro Ser Gly Asn Leu Ala Arg Asn Ser Ser Phe Ser Gln
Gln Gln1365 1370 1375Phe Ala His Gln Gly
Asn Pro Ala Val Tyr Ser Met Val His Met Asn1380 1385
1390Gly Ser Ser Gly His Met Gly Gln Met Asn Met Asn Pro Met Pro
Met1395 1400 1405Ser Gly Met Pro Met Gly
Pro Asp Gln Lys Tyr Cys1410 1415
1420522DNAArtificial SequenceDescription of Artificial SequencePRIMER
N8F1 5tcatcacttc cgacaacaga gg
22620DNAArtificial SequenceDescription of Artificial Sequence forward
primer designed from the 5' sequence of pCMVSPORT-B11, PM-U2
6ccagaaacgt cactatcaag
20719DNAArtificial SequenceDescription of Artificial Sequence reverse
primer designed from the 5' sequence of pCMVSPORT-B11, PM-U2
7ttactggaac ccccatacc
198951PRTHomo sapiens 8Cys Ile Gln Arg Phe Phe Ser Leu Asn Asp Gly Gln
Ser Trp Ser Gln1 5 10
15Lys Arg His Tyr Gln Glu Ala Tyr Leu Asn Gly His Ala Glu Thr Pro20
25 30Val Tyr Arg Phe Ser Leu Ala Asp Gly Thr
Ile Val Thr Ala Gln Thr35 40 45Lys Ser
Lys Leu Phe Arg Asn Pro Val Thr Asn Asp Arg His Gly Phe50
55 60Val Ser Thr His Phe Leu Gln Arg Glu Gln Asn Gly
Tyr Arg Pro Asn65 70 75
80Pro Asn Pro Val Gly Gln Gly Ile Arg Pro Pro Met Ala Gly Cys Asn85
90 95Ser Ser Val Gly Gly Met Ser Met Ser Pro
Asn Gln Gly Leu Gln Met100 105 110Pro Ser
Ser Arg Ala Tyr Gly Leu Ala Asp Pro Ser Thr Thr Gly Gln115
120 125Met Ser Gly Ala Arg Tyr Gly Gly Ser Ser Asn Ile
Ala Ser Leu Thr130 135 140Pro Gly Pro Gly
Met Gln Ser Pro Ser Ser Tyr Gln Asn Asn Asn Tyr145 150
155 160Gly Leu Asn Met Ser Ser Pro Pro His
Gly Ser Pro Gly Leu Ala Pro165 170 175Asn
Gln Gln Asn Ile Met Ile Ser Pro Arg Asn Arg Gly Ser Pro Lys180
185 190Ile Ala Ser His Gln Phe Ser Pro Val Ala Gly
Val His Ser Pro Met195 200 205Ala Ser Ser
Gly Asn Thr Gly Asn His Ser Phe Ser Ser Ser Ser Leu210
215 220Ser Ala Leu Gln Ala Ile Ser Glu Gly Val Gly Thr
Ser Leu Leu Ser225 230 235
240Thr Leu Ser Ser Pro Gly Pro Lys Leu Asp Asn Ser Pro Asn Met Asn245
250 255Ile Thr Gln Pro Ser Lys Val Ser Asn
Gln Asp Ser Lys Ser Pro Leu260 265 270Gly
Phe Tyr Cys Asp Gln Asn Pro Val Glu Ser Ser Met Cys Gln Ser275
280 285Asn Ser Arg Asp His Leu Ser Asp Lys Glu Ser
Lys Glu Ser Ser Val290 295 300Glu Gly Ala
Glu Asn Gln Arg Gly Pro Leu Glu Ser Lys Gly His Lys305
310 315 320Lys Leu Leu Gln Leu Leu Thr
Cys Ser Ser Asp Asp Arg Gly His Ser325 330
335Ser Leu Thr Asn Ser Pro Leu Asp Ser Ser Cys Lys Glu Ser Ser Val340
345 350Ser Val Thr Ser Pro Ser Gly Val Ser
Ser Ser Thr Ser Gly Gly Val355 360 365Ser
Ser Thr Ser Asn Met His Gly Ser Leu Leu Gln Glu Lys His Arg370
375 380Ile Leu His Lys Leu Leu Gln Asn Gly Asn Ser
Pro Ala Glu Val Ala385 390 395
400Lys Ile Thr Ala Glu Ala Thr Gly Lys Asp Thr Ser Ser Ile Thr
Ser405 410 415Cys Gly Asp Gly Asn Val Val
Lys Gln Glu Gln Leu Ser Pro Lys Lys420 425
430Lys Glu Asn Asn Ala Leu Leu Arg Tyr Leu Leu Asp Arg Asp Asp Pro435
440 445Ser Asp Ala Leu Ser Lys Glu Leu Gln
Pro Gln Val Glu Gly Val Asp450 455 460Asn
Lys Met Ser Gln Cys Thr Ser Ser Thr Ile Pro Ser Ser Ser Gln465
470 475 480Glu Lys Asp Pro Lys Ile
Lys Thr Glu Thr Ser Glu Glu Gly Ser Gly485 490
495Asp Leu Asp Asn Leu Asp Ala Ile Leu Gly Asp Leu Thr Ser Ser
Asp500 505 510Phe Tyr Asn Asn Ser Ile Ser
Ser Asn Gly Ser His Leu Gly Thr Lys515 520
525Gln Gln Val Phe Gln Gly Thr Asn Ser Leu Gly Leu Lys Ser Ser Gln530
535 540Ser Val Gln Ser Ile Arg Pro Pro Tyr
Asn Arg Ala Val Ser Leu Asp545 550 555
560Ser Pro Val Ser Val Gly Ser Ser Pro Pro Val Lys Asn Ile
Ser Ala565 570 575Phe Pro Met Leu Pro Lys
Gln Pro Met Leu Gly Gly Asn Pro Arg Met580 585
590Met Asp Ser Gln Glu Asn Tyr Gly Ser Ser Met Gly Gly Pro Asn
Arg595 600 605Asn Val Thr Val Thr Gln Thr
Pro Ser Ser Gly Asp Trp Gly Leu Pro610 615
620Asn Ser Lys Ala Gly Arg Met Glu Pro Met Asn Ser Asn Ser Met Gly625
630 635 640Arg Pro Gly Gly
Asp Tyr Asn Thr Ser Leu Pro Arg Pro Ala Leu Gly645 650
655Gly Ser Ile Pro Thr Leu Pro Leu Arg Ser Asn Ser Ile Pro
Gly Ala660 665 670Arg Pro Val Leu Gln Gln
Gln Gln Gln Met Leu Gln Met Arg Pro Gly675 680
685Glu Ile Pro Met Gly Met Gly Ala Asn Pro Tyr Gly Gln Ala Ala
Ala690 695 700Ser Asn Gln Leu Gly Ser Trp
Pro Asp Gly Met Leu Ser Met Glu Gln705 710
715 720Val Ser His Gly Thr Gln Asn Arg Pro Leu Leu Arg
Asn Ser Leu Asp725 730 735Asp Leu Val Gly
Pro Pro Ser Asn Leu Glu Gly Gln Ser Asp Glu Arg740 745
750Ala Leu Leu Asp Gln Leu His Thr Leu Leu Ser Asn Thr Asp
Ala Thr755 760 765Gly Leu Glu Glu Ile Asp
Arg Ala Leu Gly Ile Pro Glu Leu Val Asn770 775
780Gln Gly Gln Ala Leu Glu Pro Lys Gln Asp Ala Phe Gln Gly Gln
Glu785 790 795 800Ala Ala
Val Met Met Asp Gln Lys Ala Gly Leu Tyr Gly Gln Thr Tyr805
810 815Pro Ala Gln Gly Pro Pro Met Gln Gly Gly Phe His
Leu Gln Gly Gln820 825 830Ser Pro Ser Phe
Asn Ser Met Met Asn Gln Met Asn Gln Gln Gly Asn835 840
845Phe Pro Leu Gln Gly Met His Pro Arg Ala Asn Ile Met Arg
Pro Arg850 855 860Thr Asn Thr Pro Lys Gln
Leu Arg Met Gln Leu Gln Gln Arg Leu Gln865 870
875 880Gly Gln Gln Phe Leu Asn Gln Ser Arg Gln Ala
Leu Glu Leu Lys Met885 890 895Glu Asn Pro
Thr Ala Gly Gly Ala Ala Val Met Arg Pro Met Met Gln900
905 910Pro Gln Gln Gly Phe Leu Asn Ala Gln Met Val Ala
Gln Arg Ser Arg915 920 925Glu Leu Leu Ser
His His Phe Arg Gln Gln Arg Val Ala Met Met Met930 935
940Gln Gln Gln Gln Gln Gln Gln945
95094621DNAMus musculusCDS(110)..(4318) 9ggcggcgaac ggatcaaaag aatttgctga
acagtggact ccgagatcgg taaaacgaac 60tcttccctgc ccttcctgaa cagctgtcag
ttgctgatct gtgatcagg atg agt gga 118Met Ser Gly1cta ggc gaa agc tct
ttg gat ccg ctg gcc gct gag tct cgg aaa cgc 166Leu Gly Glu Ser Ser
Leu Asp Pro Leu Ala Ala Glu Ser Arg Lys Arg5 10
15aaa ctg ccc tgt gat gcc cca gga cag ggg ctt gtc tac agt ggt gag
214Lys Leu Pro Cys Asp Ala Pro Gly Gln Gly Leu Val Tyr Ser Gly Glu20
25 30 35aag tgg cga
cgg gag cag gag agc aag tac ata gag gag ctg gca gag 262Lys Trp Arg
Arg Glu Gln Glu Ser Lys Tyr Ile Glu Glu Leu Ala Glu40 45
50ctc atc tct gca aat ctc agc gac atc gac aac ttc aat
gtc aag cca 310Leu Ile Ser Ala Asn Leu Ser Asp Ile Asp Asn Phe Asn
Val Lys Pro55 60 65gat aaa tgt gcc atc
cta aag gag aca gtg aga cag ata cgg caa ata 358Asp Lys Cys Ala Ile
Leu Lys Glu Thr Val Arg Gln Ile Arg Gln Ile70 75
80aaa gaa caa gga aaa act att tcc agt gat gat gat gtt caa aaa
gct 406Lys Glu Gln Gly Lys Thr Ile Ser Ser Asp Asp Asp Val Gln Lys
Ala85 90 95gat gtg tct tct aca ggg cag
gga gtc att gat aaa gac tct tta gga 454Asp Val Ser Ser Thr Gly Gln
Gly Val Ile Asp Lys Asp Ser Leu Gly100 105
110 115ccg ctt tta cta cag gca ctg gat ggt ttc ctg ttt
gtg gtg aat cga 502Pro Leu Leu Leu Gln Ala Leu Asp Gly Phe Leu Phe
Val Val Asn Arg120 125 130gat gga aac att
gta ttc gtg tca gaa aat gtc aca cag tat ctg cag 550Asp Gly Asn Ile
Val Phe Val Ser Glu Asn Val Thr Gln Tyr Leu Gln135 140
145tac aag cag gag gac ctg gtt aac aca agt gtc tac agc atc
tta cat 598Tyr Lys Gln Glu Asp Leu Val Asn Thr Ser Val Tyr Ser Ile
Leu His150 155 160gag caa gac cgg aag gat
ttt ctt aaa cac tta cca aaa tcc aca gtt 646Glu Gln Asp Arg Lys Asp
Phe Leu Lys His Leu Pro Lys Ser Thr Val165 170
175aat gga gtt tct tgg act aat gag aac cag aga caa aaa agc cat aca
694Asn Gly Val Ser Trp Thr Asn Glu Asn Gln Arg Gln Lys Ser His Thr180
185 190 195ttt aat tgt cgt
atg ttg atg aaa aca cac gac att ttg gaa gac gtg 742Phe Asn Cys Arg
Met Leu Met Lys Thr His Asp Ile Leu Glu Asp Val200 205
210aat gcc agt ccc gaa aca cgc cag aga tat gaa aca atg cag
tgc ttt 790Asn Ala Ser Pro Glu Thr Arg Gln Arg Tyr Glu Thr Met Gln
Cys Phe215 220 225gcc ctg tct cag cct cgc
gct atg ctg gaa gaa gga gaa gac ttg cag 838Ala Leu Ser Gln Pro Arg
Ala Met Leu Glu Glu Gly Glu Asp Leu Gln230 235
240tgc tgt atg atc tgc gtg gct cgc cgc gtg act gcg cca ttc cca tcc
886Cys Cys Met Ile Cys Val Ala Arg Arg Val Thr Ala Pro Phe Pro Ser245
250 255agt cct gag agc ttt att acc aga cat
gac ctt tcc gga aag gtt gtc 934Ser Pro Glu Ser Phe Ile Thr Arg His
Asp Leu Ser Gly Lys Val Val260 265 270
275aat ata gat aca aac tca ctt aga tct tcc atg agg cct ggc
ttt gaa 982Asn Ile Asp Thr Asn Ser Leu Arg Ser Ser Met Arg Pro Gly
Phe Glu280 285 290gac ata atc cga aga tgt
atc cag agg ttc ttc agt ctg aat gat ggg 1030Asp Ile Ile Arg Arg Cys
Ile Gln Arg Phe Phe Ser Leu Asn Asp Gly295 300
305cag tca tgg tcc cag aag cgt cac tat caa gaa gct tat gtt cat ggc
1078Gln Ser Trp Ser Gln Lys Arg His Tyr Gln Glu Ala Tyr Val His Gly310
315 320cac gca gag acc ccc gtg tat cgt ttc
tcc ttg gct gat gga act att 1126His Ala Glu Thr Pro Val Tyr Arg Phe
Ser Leu Ala Asp Gly Thr Ile325 330 335gtg
agt gcg cag aca aaa agc aaa ctc ttc cgc aat cct gta acg aat 1174Val
Ser Ala Gln Thr Lys Ser Lys Leu Phe Arg Asn Pro Val Thr Asn340
345 350 355gat cgt cac ggc ttc atc
tcg acc cac ttt ctt cag aga gaa cag aat 1222Asp Arg His Gly Phe Ile
Ser Thr His Phe Leu Gln Arg Glu Gln Asn360 365
370gga tac aga cca aac cca aat ccc gca gga caa ggc atc cga cct cct
1270Gly Tyr Arg Pro Asn Pro Asn Pro Ala Gly Gln Gly Ile Arg Pro Pro375
380 385gca gca ggg tgt ggc gtg agc atg tct
cca aat cag aat gta cag atg 1318Ala Ala Gly Cys Gly Val Ser Met Ser
Pro Asn Gln Asn Val Gln Met390 395 400atg
ggc agc cgg acc tat ggc gtg cca gac ccc agc aac aca ggg cag 1366Met
Gly Ser Arg Thr Tyr Gly Val Pro Asp Pro Ser Asn Thr Gly Gln405
410 415atg ggt gga gct agg tac ggg gct tct agt agc
gta gcc tca ctg acg 1414Met Gly Gly Ala Arg Tyr Gly Ala Ser Ser Ser
Val Ala Ser Leu Thr420 425 430
435cca gga caa agc cta cag tcg cca tct tcc tat cag aac agc agc tat
1462Pro Gly Gln Ser Leu Gln Ser Pro Ser Ser Tyr Gln Asn Ser Ser Tyr440
445 450ggg ctc agc atg agc agt ccc ccc cac
ggc agt cct ggt ctt ggt ccc 1510Gly Leu Ser Met Ser Ser Pro Pro His
Gly Ser Pro Gly Leu Gly Pro455 460 465aac
cag cag aac atc atg att tcc cct cgg aat cgt ggc agc cca aag 1558Asn
Gln Gln Asn Ile Met Ile Ser Pro Arg Asn Arg Gly Ser Pro Lys470
475 480atg gcc tcc cac cag ttc tct cct gct gca ggt
gca cac tca ccc atg 1606Met Ala Ser His Gln Phe Ser Pro Ala Ala Gly
Ala His Ser Pro Met485 490 495gga cct tct
ggc aac aca ggg agc cac agc ttt tct agc agc tcc ctc 1654Gly Pro Ser
Gly Asn Thr Gly Ser His Ser Phe Ser Ser Ser Ser Leu500
505 510 515agt gcc ttg caa gcc atc agt
gaa ggc gtg ggg acc tct ctt tta tct 1702Ser Ala Leu Gln Ala Ile Ser
Glu Gly Val Gly Thr Ser Leu Leu Ser520 525
530act ctg tcc tca cca ggc ccc aaa ctg gat aat tct ccc aat atg aat
1750Thr Leu Ser Ser Pro Gly Pro Lys Leu Asp Asn Ser Pro Asn Met Asn535
540 545ata agc cag cca agt aaa gtg agt ggt
cag gac tct aag agc ccc cta 1798Ile Ser Gln Pro Ser Lys Val Ser Gly
Gln Asp Ser Lys Ser Pro Leu550 555 560ggc
tta tac tgt gaa cag aat cca gtg gag agt tca gtg tgt cag tca 1846Gly
Leu Tyr Cys Glu Gln Asn Pro Val Glu Ser Ser Val Cys Gln Ser565
570 575aac agc aga gat cac cca agt gaa aaa gaa agc
aag gag agc agt ggg 1894Asn Ser Arg Asp His Pro Ser Glu Lys Glu Ser
Lys Glu Ser Ser Gly580 585 590
595gag gtg tca gag acg ccc agg gga cct ctg gaa agc aaa ggc cac aag
1942Glu Val Ser Glu Thr Pro Arg Gly Pro Leu Glu Ser Lys Gly His Lys600
605 610aaa ctg ctg cag tta ctc acg tgc tcc
tcc gac gac cga ggc cat tcc 1990Lys Leu Leu Gln Leu Leu Thr Cys Ser
Ser Asp Asp Arg Gly His Ser615 620 625tcc
ttg acc aac tct ccc ctg gat cca aac tgc aaa gac tct tcc gtt 2038Ser
Leu Thr Asn Ser Pro Leu Asp Pro Asn Cys Lys Asp Ser Ser Val630
635 640agt gtc acc agc ccc tct gga gtg tcc tcc tca
aca tca ggg aca gtg 2086Ser Val Thr Ser Pro Ser Gly Val Ser Ser Ser
Thr Ser Gly Thr Val645 650 655tct tcc acc
tcc aat gtg cat ggg tct ctg ttg caa gag aaa cac cgg 2134Ser Ser Thr
Ser Asn Val His Gly Ser Leu Leu Gln Glu Lys His Arg660
665 670 675att ttg cac aag ttg ctg cag
aat ggc aac tcc cca gcg gag gtc gcc 2182Ile Leu His Lys Leu Leu Gln
Asn Gly Asn Ser Pro Ala Glu Val Ala680 685
690aag atc act gca gag gcc act ggg aag gac acg agc agc act gct tcc
2230Lys Ile Thr Ala Glu Ala Thr Gly Lys Asp Thr Ser Ser Thr Ala Ser695
700 705tgt gga gag ggg aca acc agg cag gag
cag ctg agt cct aag aag aag 2278Cys Gly Glu Gly Thr Thr Arg Gln Glu
Gln Leu Ser Pro Lys Lys Lys710 715 720gag
aat aat gct ctg ctt aga tac ctg ctg gac agg gat gac ccc agt 2326Glu
Asn Asn Ala Leu Leu Arg Tyr Leu Leu Asp Arg Asp Asp Pro Ser725
730 735gat gtg ctt gcc aaa gag ctg cag ccc cag gcc
gac agt ggg gac agt 2374Asp Val Leu Ala Lys Glu Leu Gln Pro Gln Ala
Asp Ser Gly Asp Ser740 745 750
755aaa ctg agt cag tgc agc tgc tcc acc aat ccc agc tct ggc caa gag
2422Lys Leu Ser Gln Cys Ser Cys Ser Thr Asn Pro Ser Ser Gly Gln Glu760
765 770aaa gac ccc aaa att aag acc gag acg
aac gag gag gta tcg gga gac 2470Lys Asp Pro Lys Ile Lys Thr Glu Thr
Asn Glu Glu Val Ser Gly Asp775 780 785ctg
gat aat cta gat gcc att ctt gga gat ttg acc agt tct gac ttc 2518Leu
Asp Asn Leu Asp Ala Ile Leu Gly Asp Leu Thr Ser Ser Asp Phe790
795 800tac aac aat cct aca aat ggc ggt cac cca ggg
gcc aaa cag cag atg 2566Tyr Asn Asn Pro Thr Asn Gly Gly His Pro Gly
Ala Lys Gln Gln Met805 810 815ttt gca gga
ccg agt tct ctg ggt ttg cga agt cca cag cct gtg cag 2614Phe Ala Gly
Pro Ser Ser Leu Gly Leu Arg Ser Pro Gln Pro Val Gln820
825 830 835tct gtt cgt cct cca tat aac
cga gcg gtg tct ctg gat agc cct gtg 2662Ser Val Arg Pro Pro Tyr Asn
Arg Ala Val Ser Leu Asp Ser Pro Val840 845
850tct gtt ggc tca ggt ccg cca gtg aag aat gtc agt gct ttc cct ggg
2710Ser Val Gly Ser Gly Pro Pro Val Lys Asn Val Ser Ala Phe Pro Gly855
860 865tta cca aaa cag ccc ata ctg gct ggg
aat cca aga atg atg gat agt 2758Leu Pro Lys Gln Pro Ile Leu Ala Gly
Asn Pro Arg Met Met Asp Ser870 875 880cag
gag aat tac ggt gcc aac atg ggc cca aac aga aat gtt cct gtg 2806Gln
Glu Asn Tyr Gly Ala Asn Met Gly Pro Asn Arg Asn Val Pro Val885
890 895aat ccg act tcc tcc ccc gga gac tgg ggc tta
gct aac tca agg gcc 2854Asn Pro Thr Ser Ser Pro Gly Asp Trp Gly Leu
Ala Asn Ser Arg Ala900 905 910
915agc aga atg gag cct ctg gca tca agt ccc ctg gga aga act gga gcc
2902Ser Arg Met Glu Pro Leu Ala Ser Ser Pro Leu Gly Arg Thr Gly Ala920
925 930gat tac agt gcc act tta ccc aga cct
gcc atg ggg ggc tct gtg cct 2950Asp Tyr Ser Ala Thr Leu Pro Arg Pro
Ala Met Gly Gly Ser Val Pro935 940 945acc
ttg cca ctt cgt tct aat cga ctg cca ggt gca aga cca tcg ttg 2998Thr
Leu Pro Leu Arg Ser Asn Arg Leu Pro Gly Ala Arg Pro Ser Leu950
955 960cag caa cag cag cag caa cag cag caa cag caa
caa caa cag cag caa 3046Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln
Gln Gln Gln Gln Gln965 970 975cag cag cag
caa cag cag cag cag caa cag cag cag atg ctt caa atg 3094Gln Gln Gln
Gln Gln Gln Gln Gln Gln Gln Gln Gln Met Leu Gln Met980
985 990 995aga act ggt gag att ccc atg
gga atg gga gtc aat ccc tat agc cca 3142Arg Thr Gly Glu Ile Pro Met
Gly Met Gly Val Asn Pro Tyr Ser Pro1000 1005
1010gca gtg ccg tct aac caa cca ggt tcc tgg cca gag ggc atg ctc tct
3190Ala Val Pro Ser Asn Gln Pro Gly Ser Trp Pro Glu Gly Met Leu Ser1015
1020 1025atg gaa caa ggt cct cac ggg tct caa
aat agg cct ctt ctt aga aac 3238Met Glu Gln Gly Pro His Gly Ser Gln
Asn Arg Pro Leu Leu Arg Asn1030 1035
1040tct ctg gat gat ctg ctt ggg cca cct tct aac gca gag ggc cag agt
3286Ser Leu Asp Asp Leu Leu Gly Pro Pro Ser Asn Ala Glu Gly Gln Ser1045
1050 1055gac gag aga gct ctg ctg gac cag ctg
cac aca ctc ctg agc aac aca 3334Asp Glu Arg Ala Leu Leu Asp Gln Leu
His Thr Leu Leu Ser Asn Thr1060 1065 1070
1075gat gcc aca ggt ctg gag gag atc gac agg gcc ttg gga att
cct gag 3382Asp Ala Thr Gly Leu Glu Glu Ile Asp Arg Ala Leu Gly Ile
Pro Glu1080 1085 1090ctc gtg aat cag gga
caa gct ttg gag tcc aaa cag gat gtt ttc caa 3430Leu Val Asn Gln Gly
Gln Ala Leu Glu Ser Lys Gln Asp Val Phe Gln1095 1100
1105ggc caa gaa gca gca gta atg atg gat cag aag gct gca cta tat
gga 3478Gly Gln Glu Ala Ala Val Met Met Asp Gln Lys Ala Ala Leu Tyr
Gly1110 1115 1120cag aca tac cca gct cag
ggt cct ccc ctt caa gga ggc ttt aac ctt 3526Gln Thr Tyr Pro Ala Gln
Gly Pro Pro Leu Gln Gly Gly Phe Asn Leu1125 1130
1135cag gga cag tca cca tcg ttt aac tct atg atg ggt cag att agc cag
3574Gln Gly Gln Ser Pro Ser Phe Asn Ser Met Met Gly Gln Ile Ser
Gln1140 1145 1150 1155caa ggc
agc ttt cct ctg caa ggc atg cat cct aga gcc ggc ctc gtg 3622Gln Gly
Ser Phe Pro Leu Gln Gly Met His Pro Arg Ala Gly Leu Val1160
1165 1170aga cca agg acc aac acc ccg aag cag ctg aga atg
cag ctt cag cag 3670Arg Pro Arg Thr Asn Thr Pro Lys Gln Leu Arg Met
Gln Leu Gln Gln1175 1180 1185agg cta cag
ggc cag cag ttt tta aat cag agc cgg cag gca ctt gaa 3718Arg Leu Gln
Gly Gln Gln Phe Leu Asn Gln Ser Arg Gln Ala Leu Glu1190
1195 1200atg aaa atg gag aac cct gct ggc act gct gtg atg
agg ccc atg atg 3766Met Lys Met Glu Asn Pro Ala Gly Thr Ala Val Met
Arg Pro Met Met1205 1210 1215ccc cag gct
ttc ttt aat gcc caa atg gct gcc cag cag aaa cga gag 3814Pro Gln Ala
Phe Phe Asn Ala Gln Met Ala Ala Gln Gln Lys Arg Glu1220
1225 1230 1235ctg atg agc cat cac ctg cag
cag cag agg atg gcg atg atg atg tca 3862Leu Met Ser His His Leu Gln
Gln Gln Arg Met Ala Met Met Met Ser1240 1245
1250caa cca cag cct cag gcc ttc agc cca cct ccc aac gtc acc gcc tcc
3910Gln Pro Gln Pro Gln Ala Phe Ser Pro Pro Pro Asn Val Thr Ala Ser1255
1260 1265ccc agc atg gac ggg gtt ttg gca ggt
tca gca atg ccg caa gcc cct 3958Pro Ser Met Asp Gly Val Leu Ala Gly
Ser Ala Met Pro Gln Ala Pro1270 1275
1280cca caa cag ttt cca tat cca gca aat tac gga atg gga caa cca cca
4006Pro Gln Gln Phe Pro Tyr Pro Ala Asn Tyr Gly Met Gly Gln Pro Pro1285
1290 1295gag cca gcc ttt ggt cga ggc tcg agt
cct ccc agt gca atg atg tca 4054Glu Pro Ala Phe Gly Arg Gly Ser Ser
Pro Pro Ser Ala Met Met Ser1300 1305 1310
1315tca aga atg ggg cct tcc cag aat gcc atg gtg cag cat cct
cag ccc 4102Ser Arg Met Gly Pro Ser Gln Asn Ala Met Val Gln His Pro
Gln Pro1320 1325 1330aca ccc atg tat cag
cct tca gat atg aag ggg tgg ccg tca ggg aac 4150Thr Pro Met Tyr Gln
Pro Ser Asp Met Lys Gly Trp Pro Ser Gly Asn1335 1340
1345ctg gcc agg aat ggc tcc ttc ccc cag cag cag ttt gct ccc cag
ggg 4198Leu Ala Arg Asn Gly Ser Phe Pro Gln Gln Gln Phe Ala Pro Gln
Gly1350 1355 1360aac cct gca gcc tac aac
atg gtg cat atg aac agc agc ggt ggg cac 4246Asn Pro Ala Ala Tyr Asn
Met Val His Met Asn Ser Ser Gly Gly His1365 1370
1375ttg gga cag atg gcc atg acc ccc atg ccc atg tct ggc atg ccc atg
4294Leu Gly Gln Met Ala Met Thr Pro Met Pro Met Ser Gly Met Pro
Met1380 1385 1390 1395ggc ccc
gat cag aaa tac tgc tga catctcccta gtgggactga ctgtacagat 4348Gly Pro
Asp Gln Lys Tyr Cys1400gacactgcac aggatcatca ggacgtggcg gcgagtcatt
gtctaagcat ccagcttgga 4408aacaaggcca gcgtgaccag cagcggggtc tgtgctgtca
tttgagcaga gctgggtctc 4468gctgaagcgc actgtctacc tgatgccctg cctctgtgtg
gcaaggtgtt ctgcctcatg 4528aggatgtgat tctggagatg gggtgttcgt aagcaccgct
ctcttacgtc actcccttct 4588gcctcgccag ccaaagtctt cacgtagatc tag
46211022DNAArtificial SequenceDescription of
Artificial Sequence forward primer A1B1/mESTF1 to screen mouse BAC
10tccttttccc agcagcagtt tg
221120DNAArtificial SequenceDescription of Artificial Sequence reverse
primer A1B1/mESTR1 used to screen mouse BAC 11atgccagaca tgggcatggg
20121402PRTMus musculus
12Met Ser Gly Leu Gly Glu Ser Ser Leu Asp Pro Leu Ala Ala Glu Ser1
5 10 15Arg Lys Arg Lys Leu Pro
Cys Asp Ala Pro Gly Gln Gly Leu Val Tyr20 25
30Ser Gly Glu Lys Trp Arg Arg Glu Gln Glu Ser Lys Tyr Ile Glu Glu35
40 45Leu Ala Glu Leu Ile Ser Ala Asn Leu
Ser Asp Ile Asp Asn Phe Asn50 55 60Val
Lys Pro Asp Lys Cys Ala Ile Leu Lys Glu Thr Val Arg Gln Ile65
70 75 80Arg Gln Ile Lys Glu Gln
Gly Lys Thr Ile Ser Ser Asp Asp Asp Val85 90
95Gln Lys Ala Asp Val Ser Ser Thr Gly Gln Gly Val Ile Asp Lys Asp100
105 110Ser Leu Gly Pro Leu Leu Leu Gln
Ala Leu Asp Gly Phe Leu Phe Val115 120
125Val Asn Arg Asp Gly Asn Ile Val Phe Val Ser Glu Asn Val Thr Gln130
135 140Tyr Leu Gln Tyr Lys Gln Glu Asp Leu
Val Asn Thr Ser Val Tyr Ser145 150 155
160Ile Leu His Glu Gln Asp Arg Lys Asp Phe Leu Lys His Leu
Pro Lys165 170 175Ser Thr Val Asn Gly Val
Ser Trp Thr Asn Glu Asn Gln Arg Gln Lys180 185
190Ser His Thr Phe Asn Cys Arg Met Leu Met Lys Thr His Asp Ile
Leu195 200 205Glu Asp Val Asn Ala Ser Pro
Glu Thr Arg Gln Arg Tyr Glu Thr Met210 215
220Gln Cys Phe Ala Leu Ser Gln Pro Arg Ala Met Leu Glu Glu Gly Glu225
230 235 240Asp Leu Gln Cys
Cys Met Ile Cys Val Ala Arg Arg Val Thr Ala Pro245 250
255Phe Pro Ser Ser Pro Glu Ser Phe Ile Thr Arg His Asp Leu
Ser Gly260 265 270Lys Val Val Asn Ile Asp
Thr Asn Ser Leu Arg Ser Ser Met Arg Pro275 280
285Gly Phe Glu Asp Ile Ile Arg Arg Cys Ile Gln Arg Phe Phe Ser
Leu290 295 300Asn Asp Gly Gln Ser Trp Ser
Gln Lys Arg His Tyr Gln Glu Ala Tyr305 310
315 320Val His Gly His Ala Glu Thr Pro Val Tyr Arg Phe
Ser Leu Ala Asp325 330 335Gly Thr Ile Val
Ser Ala Gln Thr Lys Ser Lys Leu Phe Arg Asn Pro340 345
350Val Thr Asn Asp Arg His Gly Phe Ile Ser Thr His Phe Leu
Gln Arg355 360 365Glu Gln Asn Gly Tyr Arg
Pro Asn Pro Asn Pro Ala Gly Gln Gly Ile370 375
380Arg Pro Pro Ala Ala Gly Cys Gly Val Ser Met Ser Pro Asn Gln
Asn385 390 395 400Val Gln
Met Met Gly Ser Arg Thr Tyr Gly Val Pro Asp Pro Ser Asn405
410 415Thr Gly Gln Met Gly Gly Ala Arg Tyr Gly Ala Ser
Ser Ser Val Ala420 425 430Ser Leu Thr Pro
Gly Gln Ser Leu Gln Ser Pro Ser Ser Tyr Gln Asn435 440
445Ser Ser Tyr Gly Leu Ser Met Ser Ser Pro Pro His Gly Ser
Pro Gly450 455 460Leu Gly Pro Asn Gln Gln
Asn Ile Met Ile Ser Pro Arg Asn Arg Gly465 470
475 480Ser Pro Lys Met Ala Ser His Gln Phe Ser Pro
Ala Ala Gly Ala His485 490 495Ser Pro Met
Gly Pro Ser Gly Asn Thr Gly Ser His Ser Phe Ser Ser500
505 510Ser Ser Leu Ser Ala Leu Gln Ala Ile Ser Glu Gly
Val Gly Thr Ser515 520 525Leu Leu Ser Thr
Leu Ser Ser Pro Gly Pro Lys Leu Asp Asn Ser Pro530 535
540Asn Met Asn Ile Ser Gln Pro Ser Lys Val Ser Gly Gln Asp
Ser Lys545 550 555 560Ser
Pro Leu Gly Leu Tyr Cys Glu Gln Asn Pro Val Glu Ser Ser Val565
570 575Cys Gln Ser Asn Ser Arg Asp His Pro Ser Glu
Lys Glu Ser Lys Glu580 585 590Ser Ser Gly
Glu Val Ser Glu Thr Pro Arg Gly Pro Leu Glu Ser Lys595
600 605Gly His Lys Lys Leu Leu Gln Leu Leu Thr Cys Ser
Ser Asp Asp Arg610 615 620Gly His Ser Ser
Leu Thr Asn Ser Pro Leu Asp Pro Asn Cys Lys Asp625 630
635 640Ser Ser Val Ser Val Thr Ser Pro Ser
Gly Val Ser Ser Ser Thr Ser645 650 655Gly
Thr Val Ser Ser Thr Ser Asn Val His Gly Ser Leu Leu Gln Glu660
665 670Lys His Arg Ile Leu His Lys Leu Leu Gln Asn
Gly Asn Ser Pro Ala675 680 685Glu Val Ala
Lys Ile Thr Ala Glu Ala Thr Gly Lys Asp Thr Ser Ser690
695 700Thr Ala Ser Cys Gly Glu Gly Thr Thr Arg Gln Glu
Gln Leu Ser Pro705 710 715
720Lys Lys Lys Glu Asn Asn Ala Leu Leu Arg Tyr Leu Leu Asp Arg Asp725
730 735Asp Pro Ser Asp Val Leu Ala Lys Glu
Leu Gln Pro Gln Ala Asp Ser740 745 750Gly
Asp Ser Lys Leu Ser Gln Cys Ser Cys Ser Thr Asn Pro Ser Ser755
760 765Gly Gln Glu Lys Asp Pro Lys Ile Lys Thr Glu
Thr Asn Glu Glu Val770 775 780Ser Gly Asp
Leu Asp Asn Leu Asp Ala Ile Leu Gly Asp Leu Thr Ser785
790 795 800Ser Asp Phe Tyr Asn Asn Pro
Thr Asn Gly Gly His Pro Gly Ala Lys805 810
815Gln Gln Met Phe Ala Gly Pro Ser Ser Leu Gly Leu Arg Ser Pro Gln820
825 830Pro Val Gln Ser Val Arg Pro Pro Tyr
Asn Arg Ala Val Ser Leu Asp835 840 845Ser
Pro Val Ser Val Gly Ser Gly Pro Pro Val Lys Asn Val Ser Ala850
855 860Phe Pro Gly Leu Pro Lys Gln Pro Ile Leu Ala
Gly Asn Pro Arg Met865 870 875
880Met Asp Ser Gln Glu Asn Tyr Gly Ala Asn Met Gly Pro Asn Arg
Asn885 890 895Val Pro Val Asn Pro Thr Ser
Ser Pro Gly Asp Trp Gly Leu Ala Asn900 905
910Ser Arg Ala Ser Arg Met Glu Pro Leu Ala Ser Ser Pro Leu Gly Arg915
920 925Thr Gly Ala Asp Tyr Ser Ala Thr Leu
Pro Arg Pro Ala Met Gly Gly930 935 940Ser
Val Pro Thr Leu Pro Leu Arg Ser Asn Arg Leu Pro Gly Ala Arg945
950 955 960Pro Ser Leu Gln Gln Gln
Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln965 970
975Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln
Met980 985 990Leu Gln Met Arg Thr Gly Glu
Ile Pro Met Gly Met Gly Val Asn Pro995 1000
1005Tyr Ser Pro Ala Val Pro Ser Asn Gln Pro Gly Ser Trp Pro Glu Gly1010
1015 1020Met Leu Ser Met Glu Gln Gly Pro His
Gly Ser Gln Asn Arg Pro Leu1025 1030 1035
1040Leu Arg Asn Ser Leu Asp Asp Leu Leu Gly Pro Pro Ser Asn
Ala Glu1045 1050 1055Gly Gln Ser Asp Glu
Arg Ala Leu Leu Asp Gln Leu His Thr Leu Leu1060 1065
1070Ser Asn Thr Asp Ala Thr Gly Leu Glu Glu Ile Asp Arg Ala Leu
Gly1075 1080 1085Ile Pro Glu Leu Val Asn
Gln Gly Gln Ala Leu Glu Ser Lys Gln Asp1090 1095
1100Val Phe Gln Gly Gln Glu Ala Ala Val Met Met Asp Gln Lys Ala
Ala1105 1110 1115 1120Leu Tyr
Gly Gln Thr Tyr Pro Ala Gln Gly Pro Pro Leu Gln Gly Gly1125
1130 1135Phe Asn Leu Gln Gly Gln Ser Pro Ser Phe Asn Ser
Met Met Gly Gln1140 1145 1150Ile Ser Gln
Gln Gly Ser Phe Pro Leu Gln Gly Met His Pro Arg Ala1155
1160 1165Gly Leu Val Arg Pro Arg Thr Asn Thr Pro Lys Gln
Leu Arg Met Gln1170 1175 1180Leu Gln Gln
Arg Leu Gln Gly Gln Gln Phe Leu Asn Gln Ser Arg Gln1185
1190 1195 1200Ala Leu Glu Met Lys Met Glu
Asn Pro Ala Gly Thr Ala Val Met Arg1205 1210
1215Pro Met Met Pro Gln Ala Phe Phe Asn Ala Gln Met Ala Ala Gln Gln1220
1225 1230Lys Arg Glu Leu Met Ser His His Leu
Gln Gln Gln Arg Met Ala Met1235 1240
1245Met Met Ser Gln Pro Gln Pro Gln Ala Phe Ser Pro Pro Pro Asn Val1250
1255 1260Thr Ala Ser Pro Ser Met Asp Gly Val
Leu Ala Gly Ser Ala Met Pro1265 1270 1275
1280Gln Ala Pro Pro Gln Gln Phe Pro Tyr Pro Ala Asn Tyr Gly
Met Gly1285 1290 1295Gln Pro Pro Glu Pro
Ala Phe Gly Arg Gly Ser Ser Pro Pro Ser Ala1300 1305
1310Met Met Ser Ser Arg Met Gly Pro Ser Gln Asn Ala Met Val Gln
His1315 1320 1325Pro Gln Pro Thr Pro Met
Tyr Gln Pro Ser Asp Met Lys Gly Trp Pro1330 1335
1340Ser Gly Asn Leu Ala Arg Asn Gly Ser Phe Pro Gln Gln Gln Phe
Ala1345 1350 1355 1360Pro Gln
Gly Asn Pro Ala Ala Tyr Asn Met Val His Met Asn Ser Ser1365
1370 1375Gly Gly His Leu Gly Gln Met Ala Met Thr Pro Met
Pro Met Ser Gly1380 1385 1390Met Pro Met
Gly Pro Asp Gln Lys Tyr Cys1395 1400
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic: